1
Introduction
Biomedical researchers generate, collect, and store more research data than ever. They do so in an environment that requires increasing levels of data openness and sharing as well as ever greater attention to the privacy of those from whom the data derive. Preserving those data in discoverable and accessible ways is increasingly important; however, it is no longer reasonable to collect data with the expectation that they will be stored “forever” at no cost. Data often may be worth preserving only if they have been integrated and aggregated with other related data. Resource constraints make it necessary for researchers and data archivists to consider the long-term disposition of data, data sets, and data streams, as well as to consider the long-term costs for preserving, archiving, and promoting access to those data. In many, if not most, cases, responsibility for data management shifts to different individuals and institutions as the data are collected, analyzed, curated, archived, and potentially reused for other purposes.
Given resource constraints, researchers will need to be able to estimate the quantity of data that they will collect over the course of a project as well as determine the likelihood and feasibility of data reuse when the original project has ended. Archivists might need to consider what archival principles might be imposed on the data as well as the ramifications of those principles. Archivists might also need to determine whether greater value should be placed on particular data types and to consider how risk management might inform data-preservation and archiving strategies. To support the scientific endeavor, all involved in managing the data throughout the data life cycle need to consider how choices regarding their data affect the costs of future preservation, management, and use, regardless of who bears those costs. Their decisions will require information about costs for curation and storage, revenue prospects associated with data generation or future data use, and estimated value of retained data (or, alternatively, the cost of replacing the data). While information alone will not result in the internalization of costs incurred elsewhere in the data life cycle, attention to and quantification of such costs will facilitate better allocation of resources and planning by those charged with guiding and investing in the production of scientific knowledge. Some of those costs may or may not be expressed in monetary units. Market valuation may place more value on, for example, clinical versus preclinical data in some contexts, or on one type of biomedical data versus another. Such valuations may not align with the priorities and mission of the organization bearing the cost. An understanding of how to make and use such valuations will inform data-preservation decisions.
The mission of the National Institutes of Health’s (NIH’s) National Library of Medicine (NLM) is to acquire, organize, and disseminate health-related information. NLM’s strategic plan includes accelerating discovery and advancing health through data-driven research, reaching more people through enhanced dissemination and engagement (NLM, 2018). As the largest biomedical library in the world, NLM understands that meeting the needs of
the biomedical research community requires a community-wide understanding that recent technological advances in data-collection technologies and data science require commensurate increases in resources for data curation, preservation, and discoverability (NLM, 2017). Efforts have been undertaken to understand the issues related to improving data discovery and access to NIH-funded data (e.g., Read et al., 2015), and NLM wants to strengthen a research community’s capability to value future data reuse and to estimate the cost of making reuse possible, which requires tools, methods, and practices.
At the request of NLM, the National Academies of Sciences, Engineering, and Medicine (National Academies) have prepared this report, which examines and assesses approaches and considerations for forecasting costs for preserving, archiving, and promoting access to biomedical research data. The report first identifies the different data environments (called “states” in this report) in which data must be managed and the activities that take place in those environments. The report then identifies cost drivers and the states and activities in which costs are incurred. Critical decisions that might be made during each of the data states that could influence long-term costs of curation, preservation, and access to data are identified. The purpose of this report is to provide a general framework for cost-effective decision making that encourages data accessibility and reuse for researchers, data managers, data archivists, data scientists, and institutions that support platforms for biomedical research data preservation and use. The framework is not itself a cost model; rather, it provides the set of activities that must be considered in constructing a cost model. The wide range of possible activities associated with any particular biomedical data set precludes constructing a single generic model that all readers of this report could employ. Costs will be a function of the data set characteristics, the activities that will be undertaken with the data set, and the duration of those activities. Moreover, readers of this report will be addressing cost forecasts at different times in the life cycle and with different interests.
However, the cost-forecasting framework presented in this report can be adapted by anyone responsible for managing data at any point in the life cycle, and use of the framework itself is the primary recommendation in this report. Specific recommendations regarding the application of the framework are not provided because researchers, data, repository hosts, and funding institutions and their requirements vary greatly. The report does describe the kind of environment conducive to forecasting the cost of sustainable data management, but making recommendations to specific institutions about how to create those environments is beyond the scope of this report.
THE CHARGE TO THE NATIONAL ACADEMIES AND THE STUDY COMMITTEE
NLM asked the National Academies to develop and demonstrate a cost-forecasting framework and estimate potential future benefits to research. The charge to the National Academies is provided in Box 1.1. To meet its charge, the National Academies convened an ad hoc committee of experts in areas such as biomedical sciences; biomedical informatics; cybersecurity; data science; data storage, archiving, and architectures; database systems; decision making under uncertainty; economics; ethics; health care; information theory; mathematical modeling of reliability; cost forecasting; and statistics. The members were nominated by their peers and selected by the National Academies in consideration of the balance of individual expertise and to avoid any unnecessary conflict of interest and bias. Brief biographies of the committee members are included in Appendix G.
COMMITTEE INFORMATION GATHERING AND APPROACH TO ITS TASK
Given the complexity of the task, the committee employed a series of public meetings, a workshop, multiple site visits, and one-on-one communication to hear from the numerous stakeholders in the biomedical research community. These stakeholders included biomedical-science researchers at academic or nonprofit institutions; data scientists and institutional administrators within academic, private, and public sectors; data archivists; software engineers; data platform managers; and many others—all individuals who make decisions about data throughout the entire data life cycle. The committee consulted the published literature and communicated with individuals and institutions with responsibility for managing data in different formats to understand any cost-forecasting models that might be applied and those factors that influence decision making. The committee members also drew from their collective expertise and experiences, and many of the report conclusions are based on their own observations.
The committee held five meetings in Washington, D.C.; three of these meetings included open sessions in which speakers and guests were invited to respond to questions relevant to the committee’s statement of task (Box 1.1). Agendas for the committee’s open session meetings, the workshop, and site visits appear in Appendix A.
The committee’s first meeting included presentations and a panel discussion with leadership and staff at NIH and NLM. Those individuals were asked to describe NIH’s institutional priorities and primary objectives for data management, the largest cost issues encountered to meet those objectives, and the financial mechanisms employed or being considered to meet data-related goals. The second meeting included panelists with expertise in research technology, methodologies, and workflows across the data life cycle, and in research-data collaboration from the private sector. They were asked to describe their respective methods for anticipating long-term uses and costs of data management and emerging issues that could affect data management in the future. The committee’s third meeting included discussions with the director of digital preservation from the U.S. National Archives to learn about that agency’s approaches to develop budgets for data preservation and with the deputy project leader of the World Wide Computing Grid of the European Organization for Nuclear Research (CERN) to learn about lifetime data management at CERN. A principal project manager from a commercial cloud vendor discussed the changes in technologies, data volumes and types, and data uses in the near and distant future. The committee queried all of these individuals about the positive and negative developments anticipated in the next 5 to 10 years that are likely
to affect the cost of data preservation, archiving, and access. The committee’s workshop included 15 speakers and more than 50 invited guests, and focused on elements as described in Box 1.1. Workshop participants had the opportunity to discuss (1) tools and practices that NLM could use to help researchers and funders better integrate risk-management practices and considerations into data preservation, archiving, and accessing decisions; (2) methods to encourage NIH-funded researchers to consider, update, and track lifetime data costs; and (3) burdens on the academic researchers and industry staff to implement these tools, methods, and practices. The workshop was summarized in a set of proceedings (NASEM, 2020) that was released separately from the present report.
To engage in open discussion with a number of additional stakeholders and service providers, the committee made a total of 10 site visits to a variety of institutions. To maximize efficiency, the committee first identified the types of institutions with which it should engage, identified specific institutions of those types and their locations, and then identified regions where a number of such institutions could be visited within a few days. Ultimately, the committee visited institutions in San Diego, California; Washington, D.C.; Boston, Massachusetts; and Seattle, Washington. These included visits to the National Center for Microscopy and Imaging Research; the University of California, San Diego/San Diego Supercomputer Center Advanced CyberInfrastructure Development Lab; NIH; Dana-Farber Cancer Institute; Harvard Medical School; The Broad Institute; Amazon Web Services; Institute for Systems Biology; Allen Institute; and the Fred Hutchinson Cancer Research Center. The committee also met with NIH staff across institutes that run various data repositories. Ultimately, the committee members met with more than 100 individuals with different perspectives and expertise who engaged with research data at different states within the data life cycle. Visits to any number of other institutions could have informed the committee’s deliberations, but the mix of private- and public-sector data resource managers, researchers, data scientists, and service providers provided the committee the base of information it needed. Information gaps were then filled via literature searches and personal communications between committee members and external experts.
The committee found that few think about data preservation beyond the state in which the data currently exist. Only a few of the individuals with whom the committee interacted consider how their data-related decisions affect the long-term costs associated with data preservation, curation, and access. Data regarding the costs of data resource management do not seem to be collected in any systematic way to inform future efforts. Planning for the longer term seems nascent: there are few tools or community standards available to assist with long-term planning, and such planning is at best short term in nature. Planning horizons are dictated by funding streams (e.g., federal budget allocations, grant levels) and thus extend only for the life of the project, excluding post-project data-preservation issues. Given these findings and the apparent lack of suitable examples on which to base the committee’s cost-forecasting model, the committee concluded that it would be necessary to develop an original framework on which to base a forecasting model per its charge. Once the framework was developed, the committee demonstrated its use by applying it to two use cases, per the statement of task.
FEDERAL CONTEXT
Some federal agencies, such as the National Aeronautics and Space Administration and the National Oceanic and Atmospheric Administration, are already committed to data preservation and understand that proper data preservation is a complex endeavor requiring dedicated resources over the long term. Data preservation is integrated into their cultures, and the cost of long-term preservation is included when costing a project. Other agencies have traditionally attached less importance to data preservation, and increased efforts on that front may require major cultural shifts within those agencies, a challenge that should not be underestimated. Regardless, planning horizons for agencies are often short given annual budget appropriations and that there may be legal prohibitions to planning beyond the appropriation period. These factors affect the way agencies fund external research.
Researchers who receive federal funds may be required or encouraged to do some level of planning for the disposition of their data at the end of their research projects, but research funding does not generally extend beyond the performance of the research and often does not cover data preservation. As a result, data-preservation activities are often minimal and may be conducted only as an afterthought to the research. Research grants provided by U.S. funding agencies, for example, are generally awarded only for 2- to 3-year performance periods, although some awards may be longer. Foreign funding agencies often fund research over longer performance periods with explicit
requirements related to data curation. For example, Canadian1,2 and European3,4 funding agencies offer different types of awards for periods ranging up to 7 years. German agencies have provided research funding for up to 12 years. Information-technology infrastructure and data management are explicitly incorporated into these grants.
BIOMEDICAL DATA LANDSCAPE
To provide some context for its work, the committee presents an overview of some of the current infrastructure and stakeholders that comprise the biomedical landscape, focusing mostly on infrastructure funded by NIH. The biomedical data landscape is diverse, distributed, and dynamic, characterized by an array of data repositories, databases, and platforms, which host data and make them available for reuse. These repositories are the database infrastructures where long-term stewardship, preservation, and access to research data are made possible. For the purposes of this report, the committee uses the terms “data repository” and “data archive” to refer to data infrastructure that host primary research data rather than to refer to knowledge bases that extract and aggregate analyzed data from the scientific literature. A similar distinction is adopted in the NIH Strategic Plan for Data Science (NIH, 2018). Examples of primary research data repositories include the Protein Data Bank,5 the National Institute of Mental Health Data Archive, and the National Archive of Computerized Data on Aging;6 examples of knowledge bases include UniProt7 and the Monarch Initiative.8 The distinction between a database and a knowledge base is not always clean—many digital repositories serve dual purposes—but NIH defines the primary function of a data repository to “ingest, archive, preserve, manage, distribute, and make accessible the data related to a particular system or systems.”9 The primary function of a knowledge base, according to NIH, is to “extract, accumulate, organize, annotate, and link growing bodies of information related to core datasets.”10 A third type of digital artifact that does not fit neatly into these categories is the many digital spatial atlases that cover structures such as the nervous system (e.g., Brainspan Atlas of the Developing Human,11 Cell Atlas of Mouse Brain-Spinal Cord Connectome12), urogenital system, heart, and other organs (e.g., Genito-Urinary Molecular Anatomy Project).13 An important component of biomedical data and knowledge resources is that they are usually not simple platforms for hosting data; they also comprise many software tools and services that make the data and knowledge usable and useful. Consideration of challenges in hosting physical samples (i.e., biospecimen or biosample repositories) is beyond the scope of this report.
The biomedical repository landscape spans accessible data repositories hosted by government agencies, national laboratories, research consortia, institutions and hospitals, patient advocacy organizations, researchers, journals, and commercial entities, including consortia of study sponsors. The exact number of these repositories is difficult to estimate. re3data,14 a registry of research data repositories, returns a list of 483 for the search term “basic biological and medical research,” of which 38 are in the United States. The California Digital Library, a digital research library founded by the University of California, lists more than 600 biomedical source databases
___________________
1 Information regarding the Canadian Social Sciences and Humanities Research Council is found at https://www.sshrc-crsh.gc.ca/funding-financement/programs-programmes/partnership_development_grants-subventions_partenariat_developpement-eng.aspx, accessed May 12, 2020.
2 Information regarding Canada’s Partnership Grants (stage 1) is found at https://www.sshrc-crsh.gc.ca/funding-financement/programs-programmes/partnership_grants_stage1-subventions_partenariat_etape1-eng.aspx, accessed May 12, 2020.
3 Information regarding the European Research Council (ERC) consolidator grants is found at https://erc.europa.eu/funding/consolidator-grants, accessed May 12, 2020.
4 Information regarding the ERC synergy grants is found at https://erc.europa.eu/funding/synergy-grants, accessed May 12, 2020.
5 The website for the Worldwide Protein Data Bank is https://www.wwpdb.org/, accessed December 2, 2019.
6 The website for the National Archive of Computerized Data on Aging is https://www.icpsr.umich.edu/icpsrweb/NACDA/, accessed January 2, 2020.
7 The website for UniProt is https://www.uniprot.org/, accessed December 2, 2019.
8 The website for the Monarch Initiative is https://monarchinitiative.org/, accessed December 2, 2019.
9 For example, see https://grants.nih.gov/grants/guide/pa-files/PAR-20-089.html, accessed February 20, 2020.
10 For example, see https://grants.nih.gov/grants/guide/pa-files/PAR-20-097.html, accessed February 20, 2020.
11 The website for the Brainspan Atlas of the Developing Human is http://brainspan.org/, accessed December 2, 2019.
12 The website for the Cell Atlas of Mouse Brain-Spinal Cord Connectome project is https://projectreporter.nih.gov/project_info_description.cfm?aid=9583948&icde=0, accessed December 2, 2019.
13 The website for the Genito-Urinary Molecular Anatomy Project is https://www.gudmap.org/, accessed December 2, 2019.
14 The website for the re3data registry of research data repositories is https://www.re3data.org/, accessed May 12, 2020.
(Wimalaratne et al., 2018). An analysis by the ELIXIR project lists more than 500 data resources available in Europe (Durinx et al., 2017). NLM currently lists 82 repositories on the NLM Data Sharing Repositories page.15 The Neuroscience Information Framework (NIF) has maintained a registry for online biomedical data resources since 2006. NIF lists more than 200 such primary research data repositories serving biomedicine. Approximately 120 of these repositories explicitly list support from NIH, although information was not available or recorded for many. The total number of data resources, which includes data repositories, databases, knowledge bases, and atlases in NIF, is greater than 6,000.16
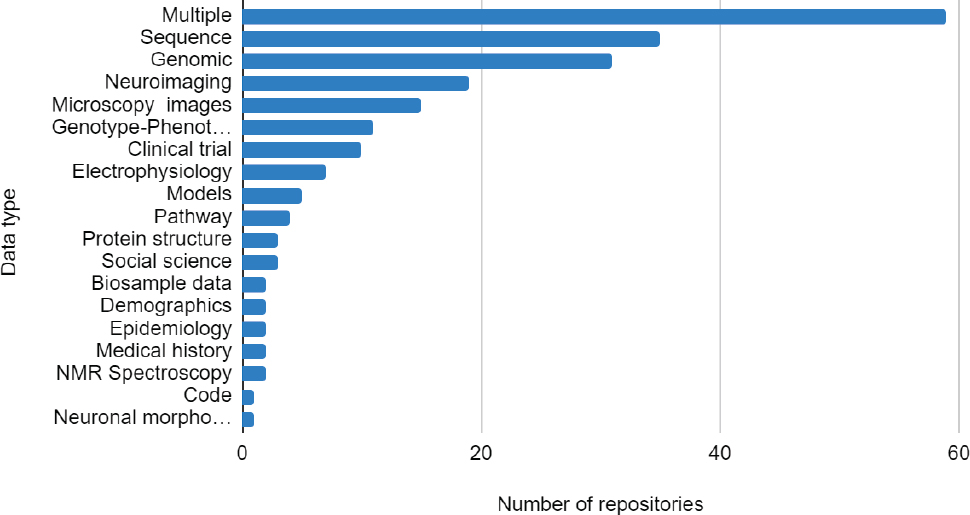
While some institutes are more invested in data repositories than others, as shown in Figure 1.1, the majority of NIH institutes and centers have funded one or more repositories. Reflecting the diversity of NIH institutes and other funding sources, the repositories cover a wide range of data types (Figure 1.2) and topics. At the same time, many generalist repositories exist that are hosted by institutions, nonprofits, and commercial entities that host data of all types, often deposited in the context of a published scientific paper. Some repositories, mostly institutional repositories and data centers supporting various research consortia, restrict data deposition, and sometimes access, to specific constituents.
Researchers generally have a choice as to where they store their data, whether the data are for private or public use. Many researchers have access to data repositories within their home institutions, or they can take advantage of specialist or generalist community repositories. In the absence of specific requirements coming from the funder or journal, the general recommendation from community organizations promoting data sharing is to use a community data repository specialized for a particular type of data (see OpenAIRE, 2019; e.g., protein-structure data might be deposited in the Protein Data Bank,17 microarray data might be deposited in the Gene Expression Omnibus). Specialist repositories generally enforce community standards and have software available to help researchers comply with these standards. They also generally provide visualization and analysis tools that work with these specialized data types (e.g., the National Center for Biotechnology Information [NCBI] Basic Local Alignment Search Tool [BLAST]18).
Most repositories do not charge the depositor a fee for submitting or hosting data (although this may be true only up to a specified size limit). For example, the Dryad Digital Repository19 is operated by a not-for-profit organization and originally hosted data associated primarily with Earth-science publications. It now functions as a cross-disciplinary data repository that is integrated as part of the manuscript submission process for more than 1,000 journals, many in biomedicine. To offset costs, Dryad charges $120 for data deposition for data sets smaller than 20 gigabytes (GB). Depositors are charged $50 for each additional 10 GB.20
Much of the ecosystem of data repositories described above has been designed to share data with third parties for the purposes of transparency and reuse. However, not all data hosted by these repositories are open—that is, made available for anyone to use and distribute. A consortium data center, for example, may make the data available only to others in the consortium, and even open repositories may have requirements (e.g., they may require approval by the Institutional Review Board that governs data use). Institutional repositories, many of which are hosted by research libraries, generally provide services for private management or public sharing of research data only for researchers within their home institutions. Not all published data are hosted within a repository. Many journals allow authors to publish small data sets that live on the journal website as supplemental materials to their papers.
The biomedical data landscape is dynamic in that new data are constantly being generated, and new data infrastructures are continually coming into existence while older ones may migrate, merge, grow stale, or be taken down. The ELIXIR project of the European Union has defined different phases of a data repository—developing, mature, and legacy—and provides some characteristics of each phase (Durinx et al., 2017). The legacy phase refers to the state in which the repository is still online but no longer growing. The NIF project shows that, of the 200 data repositories listed, only 18 have gone out of service or have merged with other entities, and approximately 10 others
___________________
15 The website for NIH’s Data Sharing Repositories is https://www.nlm.nih.gov/NIHbmic/nih_data_sharing_repositories.html, accessed August 13, 2019.
16 The website for the Neuroscience Information Framework is https://neuinfo.org/, accessed December 2, 2019.
17 The website for the Protein Data Bank is http://www.rcsb.org/, accessed December 2, 2019.
18 The website for the NCBI BLAST is https://blast.ncbi.nlm.nih.gov/Blast.cgi, accessed December 2, 2019.
19 The website for the Dryad Digital Repository is https://datadryad.org/, accessed December 2, 2019.
20 The website showing Dryad’s data publishing charges is https://datadryad.org/stash/publishing_charges, accessed May 27, 2020.

remain online but appear to be no longer actively maintained. At the same time, more data repositories are being created. A search of the NIH Research Portfolio Online Reporting Tools21 (NIH RePORTER), an online database of NIH-awarded grants, identifies another 128 data centers or data coordinating centers funded in 2018-2019 to support individual projects or consortia, many of which may be too nascent to be listed in repository catalogues.
Little information is available about what happens to data from resources that have been decommissioned. Occasionally, a resource is taken down and a plan for disposition and continued access to the data for a period of time is displayed on the website—for example, the Beta Cell Biology Consortium22 (BCBC), a project funded by the National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK). The BCBC data were transferred to the NIDDK Information Network23 before the resource went out of commission.
Less understood and studied are the practices of researchers and institutions for managing their research data before they are made available in a repository or how much such repositories and services are used. Although data management practices in the laboratory are at the front line of eventual data sharing and long-term data access, there is often a lack of incentive for researchers to think about long-term curation and preservation needs, as they do not recognize a personal benefit (see Box 1.2). The policies around data stewardship and retention at universities may not have kept pace with the digital-data revolution. Although NIH and the National Science Foundation require researchers to have data management plans specified in their grant proposals, there are no requirements for how those plans are to be formulated (see Appendix B). The committee was not able to locate any research on how much of the data generated by researchers are transferred to a more stable entity for long-term stewardship, although funder mandates and requirements are assumed to play a role in populating public archives.
The variety of expertise and types of infrastructures and services required to work with diverse data make it unlikely that biomedicine will ever be served by a single large data resource; multiple archives and data repositories will continue to exist, even for the same type of data. The advantages of such an approach are specialized tools and services and a certain amount of robustness and innovation in the ecosystem. If these repositories use the same standards, federated search across them becomes possible. Nevertheless, multiple repositories impose a cost in that separate infrastructures, staff, and tools must be maintained at each site and may, in some cases, result in less value, and less data discovery, than might otherwise accrue from a more unified resource, particularly if different standards and formats are imposed by different repositories. Therefore, after a period of innovation and separate development, it may make sense to consolidate some resources. Merging of two active data resources will lead to costs of data transfer, harmonization of data, and adaptation of technologies.
In an effort to reduce redundancy and increase functionality, the Alliance of Genome Resources24 has been formed as part of the NIH Strategic Plan for Data Science. The goals of the Alliance are to “establish a common infrastructure and software platform for data from all the [Model Organism Databases]; adopt updated data management practices; better integrate content, software, and user interfaces; improve interoperability; exchange best practices; and reduce redundancies of operation and maintenance.”25
In summary, the diversity and dynamism of data repositories and other infrastructures and the “black hole” of dark data (i.e., unpublished data; see Box 1.3) or data that are otherwise discoverable (e.g., Read et al., 2015) makes a true landscape analysis difficult. As indicated above, reliable data on the number and locations of such repositories, particularly institutional repositories, are difficult to determine,26 and few may be set up for large volumes of data. Thus, a complete accounting of biomedical data infrastructures and their content is not possible, even for publicly accessible data. To date, there is no equivalent of PubMed for biomedical data sets, although
___________________
21 The website for the NIH RePORTER is https://projectreporter.nih.gov/reporter.cfm, accessed December 2, 2019.
22 The website for BCBC is https://www.betacell.org/, accessed December 2, 2019.
23 The website for the NIDDK Information Network is https://dknet.org/, accessed December 2, 2019.
24 The website for the Alliance of Genome Resources is https://www.alliancegenome.org/, accessed December 2, 2019.
25 The website for the National Human Genome Research Institute is https://www.genome.gov/Funded-Programs-Projects/Computational-Genomics-and-Data-Science-Program/The-Alliance, accessed May 12, 2020.
26 University research data policies provide little guidance. See, for example, https://doresearch.stanford.edu/research-scholarship/research-data, https://research.columbia.edu/research-data-columbia, https://libraries.mit.edu/data-management/, http://guides.library.jhu.edu/c.php?g=813898&p=6281112, and https://ogc.umich.edu/frequently-asked-questions/research/, accessed December 12, 2019.
there have been several efforts launched by NIH and others to construct them (e.g., DataMed;27Chen et al., 2018). None of these has been fully populated.
FAIR DATA
The research community has designed a set of principles to assist reuse of data by third parties (i.e., by those other than the data producers) and drive science discovery. The findable, accessible, interoperable, and reusable (FAIR) data principles offer guidance on how to design data and data systems to make data more reusable by both humans and machines (Wilkinson et al., 2016). These principles have seen rapid endorsement by funders in both Europe and the United States. Although the principles themselves are not recommendations for implementation, they do lay out a set of 15 attributes (see Box 1.4).
Communities attempting to implement FAIR principles recognize associated costs accrued by both the data provider and those providing data access. Some costs are short lived, but others are recurrent and long lasting. Some obligations imposed by FAIR can even be viewed in perpetuity—for example, the requirement that (meta) data be assigned persistent identifiers (e.g., Digital Object Identifiers). There are short-term costs associated with providing these identifiers and then long-term costs associated with ensuring that links between the identifier and
___________________
27 The website for DataMed is https://datamed.org/, accessed December 2, 2019.
data are maintained. Similarly, the FAIR requirement that metadata are accessible, even when the data are no longer available, represents a small cost for an individual data set. However, in aggregate, the requirement imposes a societal obligation on ensuring that there are entities to maintain access to these metadata in perpetuity. Thus, implementing FAIR principles requires both technical infrastructure and organizational infrastructure. As data are transferred across entities and moved across the different stages of maturity, these services must be maintained.
On the other hand, although perhaps too early to tell, implementation of the FAIR principles also has the potential to contribute to long-term data sustainability, as agreement on and adherence to standards and best practices can conceivably lower the cost of porting data from one archive to another.
REPORT ORGANIZATION
The statement of task requests a general cost-forecasting framework that is applicable to all data resources and throughout the data life cycle. It also asks the committee to evaluate an array of considerations. To provide the basis for forecasting long-term costs for preserving, archiving, and accessing various types of biomedical data, Chapter 2 explores the three states in the data life cycle and their associated activities: (State 1) the primary
research and data management environment; (State 2) an active repository and platform where data may be acquired, curated, aggregated, accessed, and analyzed; and (State 3) a long-term preservation platform. Chapter 3 describes the economics of cost forecasting. Chapter 4 presents the cost-forecasting framework and highlights the important cost drivers, the decisions about which may affect costs throughout the data life cycle. In Chapters 5 and 6, the committee demonstrates the application of the cost-forecasting framework in biomedical contexts. Chapter 5 applies the framework of the cost forecast for a new repository and platform for biomedical research data, and Chapter 6 applies the framework to forecasting costs for new research in the primary research environment. Chapter 7 discusses potential economic, technology, policy, and legal disruptors that could affect data costs in the future. In Chapter 8, the committee offers a set of strategies, actions, and needed advances that would foster an environment conducive to responsible long-term data management decisions and cost forecasting.
The cost-forecasting framework itself is not an instrument for calculating the dollars necessary to develop or manage an information resource, but rather it is a framework that assists the cost forecaster in developing his own instrument. Each application of the framework will be unique depending on the nature of the information resource, its users and contributors, available resources, and the point of view of the forecaster. Box 1.5 outlines the major elements of the framework found throughout the report.
BENEFICIARIES OF THIS REPORT
Through its interactions with a variety of stakeholders, the committee determined that considering costs of preservation, archiving and allowing access to data beyond a short 1- to 2-year planning horizon is not part of common practice. Many researchers think about the disposition of their data after their primary research is complete and strive to make those data public. They may struggle, however, owing to the lack of financial and technical resources available once the performance period of the original research funding has ended. There are also researchers who considered the outcomes of their research to be journal articles rather than data and therefore put little thought into the long-term disposition of their data beyond that required as a condition of their research funding or journal policy. The responsibility of managing data is often transferred to the manager of a data-archiving platform, and managers of those platforms face new challenges associated with, for example, maneuvering within commercially available cloud storage and associated fee structures. Further, those managers may not be domain experts and may not understand how to retain the value of the data if the data are not properly standardized or documented.
While NLM commissioned this study, the cost-forecasting framework presented as part of this report is intended to be useful across multiple stakeholder groups, including the following:
- Researchers who need to estimate the costs involved in acquiring data, managing them effectively in the laboratory, and preparing them for submission to an archive;
- Graduate students and other shorter-term research staff who may not see or appreciate the long-term cost benefits of good decision making related to data collection and curation;
- Institutional officials at the researchers’ home institutions—these institutions bear significant shared and shifted operating and capital costs to maintain data infrastructure and supporting staff;
- Archive managers who need to estimate costs when determining the amount of funding required to fulfill their mission and who may need to transfer their archives to platforms receiving greater or lesser use;
- Program officers or other funding agency staff who are launching new programs and need to anticipate costs across the different stages of the project, including long-term preservation and access; and
- Data preservationists who will need to estimate the costs for long-term preservation ahead of procuring or accepting data.
Expanding this conversation among these and other stakeholders will not only advance data preservation, archiving, and access, but it will also foster rich scientific discovery.
REFERENCES
Barone, L., J. Williams, and D. Mickloset. 2017. Unmet needs for analyzing biological big data: A survey of 704 NSF principal investigators. PLOS Computational Biology 13(10):e1005755. https://doi.org/10.1371/journal.pcbi.1005755.
Borghi, J.A., and A.E. Van Gulick. 2018. Data management and sharing in neuroimaging: Practices and perceptions of MRI. PLoS ONE 13(7):e0200562. https://doi.org/10.1371/journal.pone.0200562.
Chen, X., A.E. Gururaj, B. Ozyurt, R. Liu, E. Soysal, T. Cohen, F. Tiryaki, et al. 2018. DataMed: An open source discovery index for finding biomedical data sets. Journal of the American Medical Informatics Association 25(3):300-308. https://doi.org/10.1093/jamia/ocx121.
Durinx, C., J. McEntyre, R. Appel, R. Apweiler, M. Barlow, N. Blomberg, C. Cook, et al. 2017. Identifying ELIXIR core data resources [version 2; peer review: 2 approved]. https://f1000research.com/articles/5-2422/v2.
NASEM (National Academies of Sciences, Engineering, and Medicine). 2020. Planning for Long-Term Use of Biomedical Data: Proceedings of a Workshop. Washington, D.C.: The National Academies Press.
NIH (National Institutes of Health). 2018. NIH Strategic Plan for Data Science. https://datascience.nih.gov/sites/default/files/NIH_Strategic_Plan_for_Data_Science_Final_508.pdf.
NLM (National Library of Medicine). 2017. Synopsis of A Platform for Biomedical Discovery and Data-Powered Health: Strategic Plan 2017-2027. https://www.nlm.nih.gov/pubs/plan/lrp17/NLM_StrategicReport_Synopsis_FINAL.pdf.
NLM. 2018. NLM Launches 2017-2027 Strategic Plan. https://www.nlm.nih.gov/news/NLM_Launches_2017_to_2027_Strategic_Plan.html.
OpenAIRE. 2019. Guides for Researchers: How to Select a Data Repository? https://www.openaire.eu/opendatapilot-repository-guide.
Read, K.B., J.R. Sheehan, M.F. Huerta, L.S. Knecht, J.G. Mork, B.L. Humphreys, and NIH Big Data Annotator Group. 2015. Sizing the problem of improving discovery and access to NIH-funded data: A preliminary study. PLoS One 10(7):e0132725. https://doi.org/10.1371/journal.pone.0132735.
Wilkinson, M.D., M. Dumontier, I. Jan Aalbersberg, G. Appleton, M. Axton, A. Baak, N. Blomberg, et al. 2016. The FAIR Guiding Principles for scientific data management and stewardship. Scientific Data 3:160018.
Wimalaratne, S.M., N. Juty, J. Kunze, G. Janée, J.A. McMurry, N. Beard, R. Jimenez, et al. 2018. Uniform resolution of compact identifiers for biomedical data. Scientific Data 5:180029.














