6
Improving Reproducibility and Replicability
STRENGTHENING RESEARCH PRACTICES: BROAD EFFORTS AND RESPONSIBILITIES
Improving substandard research practices—including poor study design, failure to report details, and inadequate data analysis—has the potential to improve reproducibility and replicability by ensuring that research is more rigorous, thoughtful, and dependable. Rigorous research practices were important long before reproducibility and replicability emerged as notable issues in science, but the recent emphasis on transparency in research has brought new attention to these issues. Broad efforts to improve research practices through education and stronger standards are a response to changes in the environment and practice of science, such as the near
ubiquity of advanced computation and the globalization of research capabilities and collaborations.
The recommendations below to improve reproducibility and replicability are generally phrased to allow flexibility in their adoption by funding agencies and the National Science Foundation (NSF). The committee’s philosophy behind this approach is that we do not apprehend all the priorities that apply to an agency such as NSF. As the committee is generally averse to displacing funds that could be applied to discovery research, we have chosen to frame our funding recommendations in terms that urge their consideration and respect the agency’s or organization’s responsibility to weigh the merits against other priorities.
In 1989, the National Academy of Sciences published its first guide to responsible research, On Being a Scientist. This booklet, directed at students in the early phases of their research careers, noted that scientific research involves making difficult decisions based on “value-laden judgments, personal desires, and even a researcher’s personality and style” (p. 1). The guide, along with updates in 1995 and 2009, laid out standards for responsible research conduct that apply across scientific fields and types of research. The most recent guide (National Academy of Sciences, National Academy of Engineering, and Institute of Medicine, 2009) describes three overarching obligations for scientists. First, because the scientific advances of tomorrow are built on the research of today, researchers have an obligation to conduct responsible research that is valid and worth the trust of their colleagues. Second, researchers have an obligation to themselves to act in a responsible and honest way. Third, researchers have an obligation to act in ways that serve the public for many reasons: research is often supported by taxpayer dollars, research results are used to inform medical and health decisions that affect people, and results underlie public policies that shape our world.
In theory, improving research practices is noncontroversial; in practice, any new requirements or standards may be a burden on tight budgets. For example, in hypothesis-testing inquiries, good research practices include conducting studies that are designed with adequate statistical power to increase the likelihood of finding an effect when the effect exists. This practice involves collecting more and better observations (i.e., reducing sampling error by increasing sample size and reducing measurement error by improving measurement precision and reliability). Although desirable in principle, this practice can involve tradeoffs in how researchers allocate limited resources of time, money, and access to limited participant populations. For example, should a researcher allocate all of these resources to test one important hypothesis or conduct lower-powered studies to test two important hypotheses, following up on the one that looks most promising? Some researchers advocate the latter approach (Finkel et al., 2017), while others have argued for the former (Albers and Lakens, 2018; Gervais et al., 2015).
Individual scientific fields have also taken steps to improve research practices, often with an explicit aim at either reproducibility or replicability, or both. Examples include the following:
- The Association for Psychological Science introduced a new journal in 2018, Advances in Methods and Practices in Psychological Science, which features articles on best practices, statistics tutorials, and other issues related to replicability. The association also offers workshops and presentations about research practices at its annual convention.
- The Society for the Improvement of Psychological Science was formed in 2016 with the explicit aim of improving research methods and practices in psychological science.
- A report of the Federation of American Societies for Experimental Biology (2016) urges all researchers to be trained in the maintenance of experimental records and laboratory notebooks; use of precise definitions and standard nomenclature for the field or experimental model; critical review of experimental design, including variables, metrics, and data analysis; application of appropriate statistical methods; and complete and transparent reporting of results.
- The ARRIVE (Animal Research: Reporting of In Vivo Experiments) guidelines for animal research give researchers a 20-point checklist of details to include in a manuscript.1 The guidelines include information about sample size, how subjects were allocated to different groups, and very specific details about the particular strain of animals used.2
- The American Vacuum Society recently published an article highlighting reproducibility and replicability issues (Baer and Gilmore, 2018).
- The Council on Governmental Relations (2018) conducted a survey among its membership to assess what resources its member institutions provide to foster rigor and reproducibility.
Perhaps the most important group of stakeholders in science are researchers themselves. If their work is to become part of the scientific record, it must be understandable and trustworthy. If they are to believe and build on the work of others, it must also be understandable and trustworthy.
___________________
1 The ARRIVE guidelines are included in a much larger set found on the EQUATOR (Enhancing the QUAlity and Transparency Of health Research) Network. See https://www.equator-network.org/reporting-guidelines.
2 The guidelines were issued by the Centre for Replacement Refinement & Reduction of Animals in Research. See https://nc3rs.org.uk/sites/default/files/documents/Guidelines/NC3Rs%20ARRIVE%20Guidelines%202013.pdf.
Thus, researchers are key stakeholders in efforts to improve reproducibility and replicability.
RECOMMENDATION 6-1: All researchers should include a clear, specific, and complete description of how a reported result was reached. Different areas of study or types of inquiry may require different kinds of information.
Reports should include details appropriate for the type of research, including
- a clear description of all methods, instruments, materials, procedures, measurements, and other variables involved in the study;
- a clear description of the analysis of data and decisions for exclusion of some data and inclusion of other;
- for results that depend on statistical inference, a description of the analytic decisions and when these decisions were made and whether the study is exploratory or confirmatory;
- a discussion of the expected constraints on generality, such as which methodological features the authors think could be varied without affecting the result and which must remain constant;
- reporting of precision or statistical power; and
- a discussion of the uncertainty of the measurements, results, and inferences.
Education and Training
In order to conduct research that is reproducible, researchers need to understand the importance of reproducibility, replicability, and transparency, to be trained in best practices, and to know about the tools that are available. Educational institutions and others have been incorporating reproducibility in classrooms and other settings in a variety of ways. For example:
- A new course at the University of California, Berkeley, “Reproducible and Collaborative Data Science,” introduces students to “practical techniques and tools for producing statistically sound and appropriate, reproducible, and verifiable computational answers to scientific questions.”3 The university has also created the Berkeley Initiative for Transparency in the Social Sciences.4
___________________
3 For the course description, see https://berkeley-stat159-f17.github.io/stat159-f1.
4 See https://www.bitss.org.
- At New York University (NYU) and Johns Hopkins University, reproducibility modules have been added to existing computation courses.
- Librarians at NYU hold office hours for questions about reproducibility and offer tutorials, such as “Citing and Being Cited: Code and Data Edition,” which teaches students how and why to share data and code.
- Also at NYU, the Moore-Sloan Data Science Environment5 has created the Reproducible Science website6 to serve as an open directory of reproducibility resources for issues beyond computational reproducibility.
- A nonprofit organization, the Carpentries, teaches foundational coding and data science skills to researchers worldwide, offering courses such as “Software Carpentry” and “Data Carpentry.”7
- Various other entities offer various short courses.8
New tools and methods for computation and statistical analysis are being developed at a rapid pace. The use of data and computation is evolving, and the ubiquity and intensity of data are such that a competent scientist today needs a sophisticated understanding of computation and statistics. Investigators want and need to use these tools and methods, but their education and training have often not prepared them to do so. Researchers need to understand the complexity of computation and acknowledge when outside collaboration is necessary. Adequate education and training in computation and statistical analysis is an inherent part of learning how to be a scientist today.
Improving Knowledge and the Use of Statistical Significance Testing
A particular source of non-reproducibility discussed in Chapter 5 is the misunderstanding and misuse of statistical significance testing. The American Statistical Association (ASA) (2016) published six principles about p-values, noting that in its 177 years of existence, it had never previously taken a stance on a specific matter of statistical practice. However, given the recent discussion about reproducibility and replicability, the ASA (2016, p. 129) decided to release these principles in the hopes that they would “shed light on an aspect of our field that is too often misunderstood and misused in the broader research community.” The principles cover key issues in statistical reporting:
___________________
5 See http://msdse.org.
6 See https://reproduciblescience.org.
7 See https://carpentries.org.
8 For example, see http://eriqande.github.io/rep-res-web; https://o2r.info/2017/05/03/egu-short-course-recap; and https://barbagroup.github.io/essential_skills_RRC [January 2019].
- P-values can indicate how incompatible the data are with a specified statistical model.
- P-values do not measure the probability that the studied hypothesis is true, or the probability that the data were produced by random chance alone.
- Scientific conclusions and business or policy decisions should not be based only on whether a p-value passes a specific threshold.
- Proper inference requires full reporting and transparency.
- A p-value, or statistical significance, does not measure the size of an effect or the importance of a result.
- By itself, a p-value does not provide a good measure of evidence regarding a model or hypothesis.
More recently, The American Statistician, which is the official journal of the ASA, released a special edition, titled “Statistical Inference in the 21st Century: A World Beyond P < 0.05,” focused on the use of p-values and statistical significance.” In the introduction to the special edition, Wasserstein and colleagues (2019) strongly discourage the use of a statistical significance threshold in reporting results due to overuse and wide misinterpretation.
RECOMMENDATION 6-2: Academic institutions and institutions managing scientific work such as industry and the national laboratories should include training in the proper use of statistical analysis and inference. Researchers who use statistical inference analyses should learn to use them properly.
EFFORTS TO IMPROVE REPRODUCIBLITY
Chapters 4 and 5 cover current knowledge on the context, extent, and causes of, non-reproducibility and non-replicability respectively. In the case of non-reproducibility, the causes include inadequate recordkeeping and nontransparent reporting. Improving computational reproducibility involves better capturing and sharing information about the computational environment and steps required to collect, process, and analyze data. All of the sources of non-reproducibility also impair replicability efforts, since they deprive researchers the information that is useful in designing or undertaking replication studies. These primary causes of non-reproducibility also directly contribute to non-replicability in that they make errors in analysis more likely, and make it more difficult to detect error, data fabrication, and data falsification.
Recordkeeping
Researchers typically execute multiple computational steps and lines of reasoning as they develop models, perform analyses, or formulate and test hypotheses. This process may involve executing multiple computational steps and using a variety of tools, both of which may require a set of inputs (including data and parameters) and are executed in a computational environment comprised of hardware and software (e.g., operating system and libraries). The automatic capture of computational details is becoming more common across domains to aid in recordkeeping. This section reviews some of the tools that are available for that task. In the discussion below, mention specific tools or platforms to highlight a current capability; this should not be seen as an endorsement by this committee.
One example comes from physicists working at CERN, who have developed methods to “capture the structured information about the research data analysis workflows and processes to ensure the usability and longevity of results” (Chen et al., 2018, p. 119). Figure 6-1 shows how the large-scale CERN collaboration has developed infrastructure for capturing computational details to allow for reproducibility and data reuse. In other fields, open source workflow-based visualization tools, such as VisTrails, have been developed to automatically capture computational details.9
Smaller groups or individual researchers may also capture computational details necessary for reproducibility. A computer scientist may run a new simulation code in a computing cluster, copy the results to her desktop, and analyze them using an interactive notebook (i.e., additional code). To analyze additional simulation results, she can automate the complete process by creating a scientific workflow. After the results are published, a detailed provenance of the process needs to be included to enable others to reproduce and extend them. This information includes the description of the data, the computational steps followed, and information about the computational environment.
For a paper that investigates Galois conjugates of quantum double models (Freedman et al., 2011), each figure is accompanied by its provenance, consisting of the workflow used to derive the plot, the underlying libraries invoked by the workflow, and links to the input data—that is, simulation results stored in an archival site (see Figure 6-2). This provenance information allows all results in the paper to be reproduced. In the
___________________
9 Traditional workflow systems are used to automate repetitive computations, very much like a script. For example, instead of typing commands on a shell, a workflow can be created that automatically issues the commands. The open source VisTrails software does this and captures the evolution of the computations (e.g., the use of different simulations, parameter explorations). See https://www.vistrails.org/index.php/Main_Page.
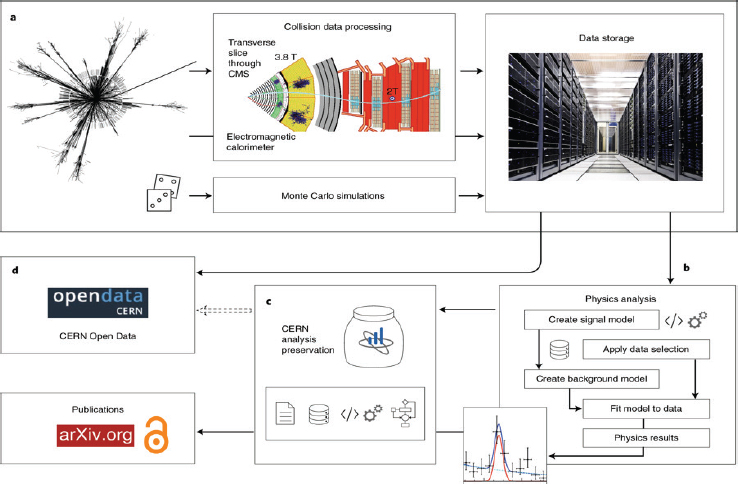
NOTES: (a) The experimental data from proton–proton collisions in the Large Hadron Collider (LHC) are collected, curated, and stored. The raw experimental data is filtered and processed to give the collision dataset formats that are suitable for physics analyses (i.e., data curation). In parallel, the computer simulations also produce data and provide necessary comparison of experimental data with theoretical predictions. (b) The stored collision and simulated data are then released for individual physics analyses across a large collaboration. A physicist may perform further data reduction and selection procedures, which are followed by a statistical analysis on the data. The analysis assets being used by the individual researcher include the information about the collision and simulated datasets, the detector conditions, the analysis code, the computational environments, and the computational workflow steps used by the researcher to derive the histograms and the final plots as they appear in publications. (c) The CERN analysis preservation service captures all the analysis assets and related documentation via a set of “push” and “pull” protocols, so that the analysis knowledge and data are preserved in a trusted long-term digital repository for preservation purposes. (d) The CERN open data service publishes selected data as they are released by the LHC collaborations into the public domain, after an embargo period of several years, depending on the collaboration data management plans and preservation policies (Chen et al., 2018, p. 114).
SOURCES: Chen et al. (2018, Fig. 1). CERN (a); Dave Gandy (b, c, code icon); Simplelcon (b, c, gear icon); Andiran Valeanu (b, c, data icon); Umar Irshad (c, paper icon); Freepik (c, workflow icon); https://www.nature.com/articles/s41567-018-0342-2#rightslink. See https//creativecommons.org/licenses/by/4.0.
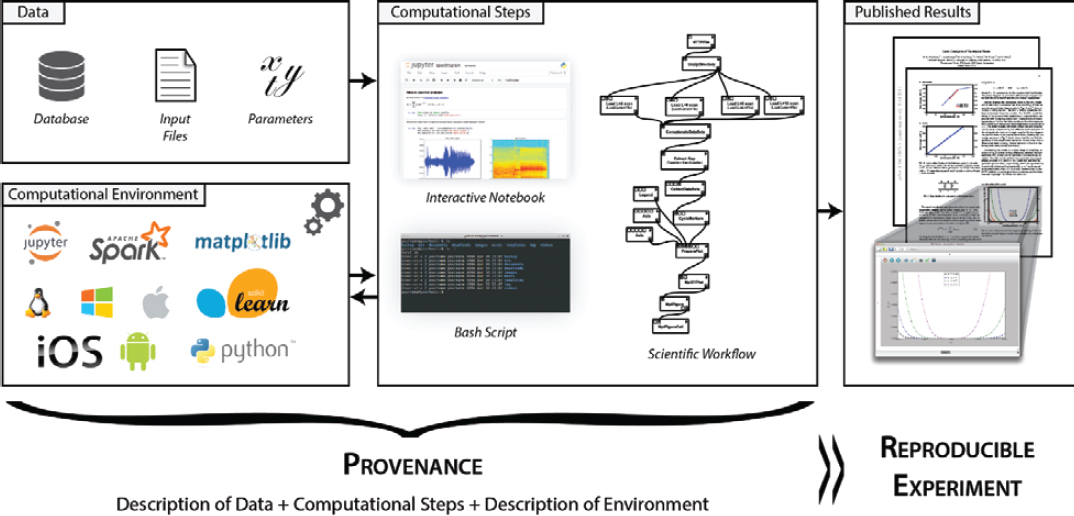
NOTE: A full description of each, or the study’s provenance, is required for reproducible research.
SOURCE: Chirigati et al. (2016).
PDF version of the paper,10 the figures are active, and when clicked on, the corresponding workflow is loaded into the VisTrails system and executed on a reader’s machine. The reader may then modify the workflow, change parameter values, and input data (Stodden et al., 2014b, p. 35).
Source Code and Data Version Control
In computational environments, several researchers may be working on shared code or data files. The changes to the code or data files affect the results. In order for another researcher to reproduce results (or even to clearly understand what was done), the version of the code or data file is an important reporting detail. However, manual recordkeeping of the multiple changes by each user (or even a single user) is burdensome and adds significant work to the research effort.
Version-control systems can automatically capture the history of all changes made to the source code of a computer program, often saved as a text file. This creates a history of changes and allows developers to better understand the code and to identify possible problems or errors.
One of the most extensively used version-control systems, Git,11 is a free and open source distributed version-control system. Scientists are increasingly adopting it as a necessary piece in the reproducibility toolbox (Wilson et al., 2014; Blischak et al., 2016). Recently, the concept of version control has been extended to data files. These files are generally too large to be stored in standard version-control systems, and they are often in a binary format that cannot be versioned. Rather, data version control excludes the large data files from the main versioned repository, automatically collects data provenance, records data processing steps into a (reproducible) pipeline, and connects with cloud services where large data files are stored.
Scientific Workflow-Management Systems
Scientific workflows represent the complex flow of data products through various steps of collection, transformation, and analysis to produce an interpretable result. Capturing provenance of the result is increasingly difficult to do using manual processes. Thus, to support computational reproducibility, efforts have been under way for several years to develop workflow-management systems that capture and store data and workflow provenance automatically. With such systems, results can be reliably linked to the computational process that derived them, and computational tasks
___________________
10 See http://arxiv.org/abs/1106.3267.
11 See https://git-scm.com.
can be automated, allowing them to be rerun and shared. We present some examples below.
In the life sciences, the Taverna project began as a tool to compose bioinformatics workflows (Oinn et al., 2004), including programmatic access to web-based data repositories and analysis tools. It is now an extensive set of open source tools that are used in biology, chemistry, meteorology, social sciences, and other fields.12
In physics, the Chimera system (Foster et al., 2002) originated in support of data-intensive physics as a means to capture and automate a complex pipeline of transformations on the data by external software. An early prototype was tested in the analysis of data from the Sloan Digital Sky Survey,13 where image and spectroscopic data are transformed in several stages to computationally locate galaxy clusters in the images. The workflow involved reading and writing millions of data files (Annis et al., 2002). Chimera enables on-demand generation of the derived data through its “virtual data” system, reducing data storage requirements, while at the same time making the transformation pipeline reproducible.
The Open Science Framework, developed by the Center for Open Science (2018), is a cloud-based project management tool that emerged as part of efforts to replicate psychological research that can be used by researchers in other fields.14 It is open source and free to use, and integrates a number of other open scientific infrastructure resources.
Bowers and Ludäscher (2005) constructed a formal model describing scientific workflows and separating concerns, such as communication or data flow and task coordination or orchestration. Modeling the basic components of scientific workflows enables them to propose various workflow design strategies, including task-, data-, structure-, and semantic-driven. The Kepler workflow-management system (Ludäscher et al., 2006), which arose in ecology and environmental communities, leverages this formal approach for creating, analyzing, and sharing complex scientific workflows.15 It can access and connect disparate data sources (e.g., streaming sensor data, satellite images, simulation output) and integrate software components for analysis or visualization. Kepler is used today in many fields, including bioinformatics, chemistry, genetics, geoinformatics, oceanography, and phylogeny. It is free and open source and has been supported over the years by various agencies, including NSF.
The previously described VisTrails system (see Callahan et al., 2006; Freire et al., 2006) goes beyond workflow management and provenance
___________________
12 See https://taverna.incubator.apache.org.
13 See https://www.sdss.org.
14 See https://osf.io.
capture by adding support for exploratory visualization. It is able to maintain a detailed history of all steps followed in the course of an exploration involving computational tasks that are iteratively modified. Thus, it captures provenance of the workflow evolution.
An NSF-funded workshop in 2006, titled “Challenges of Scientific Workflows,” focused on the questions of how to represent, capture, share, and manage workflows, and the research needs in this arena. Among the many workshop recommendations was the idea of integrating workflow representations into the scholarly record (Gil et al., 2007). A subsequent large community effort led to the specification of an Open Provenance Model (Moreau et al., 2007, 2011), which enables unambiguously sharing provenance information. This model motivated interoperable new tools and continued research and development in workflow and data provenance.16 Today, automated scientific workflows are essential in data-intensive and large-scale science missions that aim to be computationally reproducible (Deelman et al., 2018).
Tools for Reproduction of Results
After researchers capture the data and workflow provenance, and possibly use some technology to package the full computational environment used in generating some results, other researchers who want to reproduce the results in their local environment may still face challenges. A number of solutions have been proposed that attempt to simplify reproduction through the use of virtualization and cloud computing. For example, a researcher using ReproZip will create a package to share with others who can unpack, inspect, and reproduce the computational sequence in their own environment; they can use virtualization tools (such as virtual machines and containers), or they can execute the package in the cloud, without the need to install any additional software packages (Rampin et al., 2018). Virtual machines encapsulate an entire computational environment, from the operating system up through all the layers of software, including the whole file system.
Howe (2012) describes how virtual machines hosted on public clouds can enable reproducibility. Transparency can be compromised if the original researcher makes available a virtual machine containing executable software as “black boxes,” without supplying the source code. Following good practices for documenting and sharing computational artifacts, however, the combination of virtual machines and public cloud has proved valuable for reproducibility in several domains, such as microbial ecology and bioinformatics (Ragan-Kelley et al., 2013). A newer solution that is gaining
___________________
backers in the reproducible-research community are container technologies, such as Docker17 (Boettiger, 2015). Researchers can build container images that work similarly to virtual images, but instead of bundling all the data and software dependencies in a single file, the container image is built from stackable pieces. Being a more lightweight solution, containers are smaller and have less overhead than virtual images. They are being used in archaeological research (Marwick, 2017), genomics (Di Tommaso et al., 2015), phylogenomics (Waltemath and Wolkenhauer, 2016), and many other science fields. The Docker software had contributors from many organizations, including Google, IBM, and Microsoft as well as the team at Docker, Inc. It has been widely adopted in industry (Vaughan-Nichols, 2018), which led to fast innovation and sustainability that researchers can directly benefit from. The Software Sustainability Institute in the United Kingdom held a June 2017 workshop, “Docker Containers for Reproducible Research,” where talks covered applications in bioinformatics, deep learning, high-energy physics, and metagenomics, among more general technical topics.
Interactive computational notebooks are another technology supporting reproducible research (Shen, 2014). Jupyter is an open source project developing a set of tools for interactive computation and data analysis, enabling researchers to fully narrate their analysis with text and multimedia content.18 The narratives and the computational analysis are saved in a Jupyter Notebook, which can be shared with other researchers to reproduce the computations. Notebooks organize the content into cells: code cells that can be individually executed and produce output below them, and content cells written with Markdown formatting syntax. The output of code cells can be of any type, including data plots and interactive visualizations (Kluyver et al., 2016). Scientists are increasingly adopting Jupyter for their own exploratory computing, sharing knowledge within their communities, and publishing alongside traditional academic papers. One example is the publication of the confirmed detection of gravitational waves by the Laser Interferometer Gravitational-Wave Observatory (LIGO) experiment. The researchers published Jupyter notebooks that reproduced the analysis of the data displaying the signature of a binary black-hole merger.19
Recent technological advances in version control, virtualization, computational notebooks, and automatic provenance tracking have the potential to simplify reproducibility, and tools have been developed that leverage these technologies. However, given that computational reproducibility requirements vary widely even within a discipline, there are still
___________________
17 Docker is a free and open source; see https://www.docker.com/resources/what-container.
18 See https://jupyter.org.
19 See https://www.gw-openscience.org/data and https://doi.org/10.7935/K53X84K2.
many questions to be answered both to understand the gaps left by existing tools and to develop principled approaches that fill those gaps. Making the creation of reproducible experiments easy and an integral part of scientists’ computational environments would provide a great incentive for much broader adoption of reproducibility.
RECOMMENDATION 6-3: Funding agencies and organizations should consider investing in research and development of open-source, usable tools and infrastructure that support reproducibility for a broad range of studies across different domains in a seamless fashion. Concurrently, investments would be helpful in outreach to inform and train researchers on best practices and how to use these tools.
Publication Reproducibility Audits
One approach that could be taken by publishers is to assess the reproducibility of a manuscript’s results before the manuscript is published, using the data and code provided by the authors. One publication that does so is the American Journal of Political Science, which uses paid external contractors to assess reproducibility (Jacoby, 2017). The journal submits accepted manuscripts to an external reproducibility check before they are published. As discussed in Chapter 4, since the journal began this process in 2015, the external check has almost always (e.g., 108 of 116 articles) found some issue that requires the author to provide more information or make a change. This prepublication checking is an expensive and labor-intensive process, but it allows the journal to be more confident in the reproducibility of the work that is published. The external check adds about 52 days to the publishing process and, as noted above, requires an average of 8 person-hours. The files with the author-provided information are also made available to the public. Other journals submit some but not all manuscripts to such a check, sometimes during the review process and sometimes when a manuscript is accepted.
One advantage of prepublication reproducibility checks is that they encourage authors to be careful in how they conduct, report, and document their analyses. Knowing that the journal will (or may) send the data and code to be checked for reproducibility may make authors more careful to document their data and analyses clearly and to make sure the reported results are free of errors. Another important benefit of this approach is that verifying all manuscripts for reproducibility before publication should lead to a high rate of reproducibility of published results. Whether submitting some but not all manuscripts to such a check has an effect on the non-checked papers is an open question. The primary downside of this practice is that it is expensive, time consuming, and labor intensive. It is
also challenging to make sure that statistical analysis code can be executed by others who may not have the same software (or the same version of the software) as the authors.
A number of other initiatives related to publication and reporting of research results are discussed below in the context of supporting reproducibility.
RECOMMENDATION 6-4: Journals should consider ways to ensure computational reproducibility for publications that make claims based on computations, to the extent ethically and legally possible. Although ensuring such reproducibility prior to publication presents technological and practical challenges for researchers and journals, new tools might make this goal more realistic. Journals should make every reasonable effort to use these tools, make clear and enforce their transparency requirements, and increase the reproducibility of their published articles.
OVERCOMING TECHNOLOGICAL AND INFRASTRUCTURE BARRIERS TO REPRODUCIBILITY
Even if complete information about the computational environment and workflow of a study are accurately recorded, computational reproducibility is only practically possible if this information is available to other researchers. The open and persistent availability of digital artifacts, such as data and code, are also essential to replicability efforts, as explained in Chapter 5. Yet, barriers related to costs, lack of infrastructure, disciplinary culture, and weak incentives act as barriers to achieving persistent availability of these digital objects (National Academies of Sciences, Engineering, and Medicine, 2018). A number of relevant initiatives are under way to overcome the technological and infrastructure barriers, as discussed below. Initiatives on barriers related to culture and incentives are discussed later in the chapter.
Archival Repositories and Open Data Platforms
The widespread availability of repositories where researchers can deposit the digital artifacts associated with their own work, as well as find the work of others, is an enabling condition for improving reproducibility and replicability. A wide range of organizations maintain repositories, including research institutions, disciplinary bodies, and for-profit companies. The FAIR (findable, accessible, interoperable, and reusable) data principles, published in 2016 and discussed in more detail below, provide a framework for the management and stewardship of research data aimed at facilitating the sharing and use of research data.
In order to share data, code, and other digital artifacts, researchers need repositories that meet a set of standard requirements. A number of new repositories have been developed in recent years, either institutionally based or discipline specific. They can be found by means of directories of open access repositories, such as OpenDOAR (Directory of Open Access Repositories)20 and ROAR (Registry of Open Access Repositories).21
The minimum requirements for an archival repository are that it is searchable by providing a unique global identifier for the deposited artifact, has a stated guarantee of long-term preservation, and is aligned with a standard set of data access and curation principles. The details of each of these three requirements are not well established across science and engineering. Most commonly, to meet these requirements, a Digital Object Identifier (DOI, see below) is used as a unique global identifier, long-term preservation guarantees are at least 10 years, and FAIR principles are used. Box 6-1 describes two repositories that satisfy these requirements; other examples include Dataverse22 and Dryad.23
NSF has funded open science efforts in specific disciplines. In earth sciences, The Magnetics Information Consortium (MagIC) provides a data archive that allows the discovery and reuse of such data for the broader earth sciences community.24 MagIC began in 2002 as an NSF-funded project to develop a comprehensive database for archiving of paleontology and rock magnetic data, from laboratory measurements to a variety of derived data and metadata, such as the positions of the spin axis of the Earth from the point of view of the wander continents and the variations of the strength and direction of the field through time, to changes in environmentally controlled rock magnetic mineralogy. Closely linked to the MagIC project is open source software for the conversion of laboratory data to a common data format that allows interpretation of the data in a consistent and reproducible manner. Once published, the data and interpretations can be uploaded into the MagIC database. All software involved with the MagIC project is freely available on GitHub repositories. MagIC also maintains an open access textbook on rock and paleomagnetism and links the data to the original publications (only a portion of which are currently openly available).
NSF also supported the Paleo Perspectives on Climate Change (P2C2) Program, which funds much of the relevant, ongoing paleoclimate research, with the goals of generating proxy datasets that can serve as tests
___________________
20 See http://v2.sherpa.ac.uk/opendoar.
21 See http://roar.eprints.org.
22 See https://dataverse.org.
23 See https://datadryad.org.
24 See http://earthref.org/MagIC; also see National Academies of Sciences, Engineering, and Medicine (2018, p. 92).
for climate models and synthesizing proxy and model data to understand longer-term and higher-magnitude climate system variability not captured by the instrumental record.25
Code Hosting and Collaboration Platforms
Version-control systems tools like Git26 are often used in concert with hosting services like GitHub, Bitbucket, GitLab, and others. These hosting services offer “repositories” (note that this term is used differently in the
___________________
25 See https://www.nsf.gov/funding/pgm_summ.jsp?pims_id=5750.
26 See https://git-scm.com.
software world than in the data world) where authors share their code, while at the same time synchronizing the history of changes with their version-control tool, used locally in their working computers.
Increasing numbers of open source research software projects are hosted on these services, and researchers are taking advantage of them for more than code: writing reports and manuscripts, sharing supplementary materials for papers, and other artifacts. Large research organizations often create their own space on GitHub for collecting their project repositories, including the National Aeronautics and Space Administration,27 Allen Institute for Brain Science,28 and Space Telescope Science Institute,29 among others (Perez-Riverol et al., 2016). These hosting services are provided by companies—for example, GitHub is owned by Microsoft—a fact that could raise concern if researchers rely on them for code, data access, management for reproducibility, and sharing because companies can disappear or change their focus or priorities (see National Academies of Sciences, Engineering, and Medicine, 2018).
Digital Object Identifiers
A DOI is a unique sequence of characters assigned to a digital object by a registration agency, identifying the object and providing a persistent link to it on the Internet (Barba, 2019). The DOI system is an international standard, is interoperable, and has been widely adopted. Almost all scholarly publications assign a DOI to the articles they publish, and archival repositories assign them to every artifact they receive. A DOI contains standard metadata about the object, including the URL (uniform resource locator) to where the object can be found online. When the URL changes for any reason, the publisher can update the metadata so that the DOI still resolves to the object’s location.
The permanent and unambiguous identification afforded by a DOI makes a wide variety of research artifacts shareable and citable. Archival-quality repositories (e.g., Dataverse, Dryad, Figshare, Zenodo) assign a DOI to all artifact deposits, whether they are data, figures, or a snapshot of the complete archive of a research software.
Journals with a data sharing policy often require the data that a paper relied on or produced be deposited in an archival-quality repository with a DOI. For example, PLOS journals require authors to make their data available (with few exceptions): “Repositories must assign a stable persistent
___________________
27 See https://github.com/nasa.
identifier (PID) for each dataset at publication, such as a digital object identifier (DOI) or an accession number” (PLOS ONE, 2018).
Obsolescence of Data and Code Storage
More thorough reporting that includes, for example, the unique identifier to the correct version of each library and the research code that produced the results can attenuate the problem of obsolescence of data and code storage. Proper use of archival repositories also helps. Researchers increasingly adopt best practices, such as making an archival deposit of the source code associated with a publication and citing its DOI, rather than simply including a link to a GitHub (or similar) repository, which could be later deleted (Barba, 2019). As discussed above, virtual machines and containers are two available solutions to package an entire software stack, ensuring the correct versions of dependencies are available.
Sometimes, however, technological breakdowns occur. A dataset could be in a format that is no longer legible with current computers. University libraries are developing a more comprehensive role to aid researchers in the curation of digital artifacts, particularly research data, and they will increasingly participate in reproducibility initiatives. For example, Yale University Library has begun a digital preservation project that involves building infrastructure for on-demand access to old software, applying emulation technologies (Cummings, 2018).
Ensuring the longevity and openness of digital artifacts is a new challenge. Research institutions, research funders, and disciplinary groups all recognize that they have responsibilities for the long-term stewardship of digital artifacts, and they are developing strategies for meeting those needs. For example, university libraries are developing strategies for covering the associated costs (Erway and Rinehart, 2016). Although many research funders support the costs of data management during the life of the project, including preparing data for deposit in a repository, the ongoing costs of stewardship that are borne by the institution need to be covered by some combination of indirect budgets, unrestricted or purpose-directed funds, and perhaps savings from cutting other services.
Some disciplinary communities have also developed robust institutions and support structures for the stewardship of digital artifacts. One longstanding example from the social and behavioral sciences is the Inter-university Consortium for Political and Social Research (ICPSR)30. Supported by member dues as well as by public and private grants, ICPSR maintains an archive of community datasets, offers education and training programs, and performs other data-related services.
___________________
Yet there are wide disparities among research fields in their readiness to take on the tasks of preserving and maintaining access to digital artifacts. There are many discipline-specific considerations, such as which data and artifacts to store, in what formats, how best to define and establish the institutional and physical infrastructure needed, and how to set and encourage standards. The needs and requirements of governmental bodies, funding organizations, publishers, and researchers need to be taken into account. For all, an overarching concern is how much these long-term efforts will cost and how they will be supported. It is important to keep in mind that storage itself is not the only expense; there are other life-cycle costs associated with accessioning and de-accessioning data and artifacts, manually curating and managing the information, updating to new technology, migrating data and artifacts to new and changing systems, and other activities and costs that may enable the data to be used.
In the past several years, several executive and legislative actions have sought to provide incentives for data and artifact sharing and to encourage standardized processes. A 2013 memorandum from the Office of Science and Technology Policy31 directed all federal agencies with expenditures of more than $100 million to develop plans for improving access to digital data that result from federally funded research. In 2019, Congress enacted the Open, Public, Electronic, and Necessary Government Data Act,32 which requires several advancements in data storage and access, including that open government data assets are to be published as machine-readable data. In addition, it requires that “each agency shall (1) develop and maintain a comprehensive data inventory for all data assets created by or collected by the agency, and (2) designate a Chief Data Officer who shall be responsible for lifecycle data management and other specified functions.” The act also establishes in the U.S. Office of Management and Budget a Chief Data Officer Council for establishing government-wide best practices for the use, protection, dissemination, and generation of data and for promoting data sharing agreements among agencies.
Some research funders are already encouraging more openness in their funded research. For example, the National Institutes of Health (NIH) stores and makes available data and artifacts from work funded throughout all of its institutes. To help better estimate and prepare for increased data needs, NIH has recently funded a study by the National Academies of Sciences, Engineering, and Medicine to examine the long-term costs of preserving, archiving, and accessing biomedical data.
For some “big science” research projects that are computing-intensive and generate large amounts of data, management and long-term preservation
___________________
31 See https://obamawhitehouse.archives.gov/sites/default/files/microsites/ostp/ostp_public_access_memo_2013.pdf.
32 This act was part of the Foundations for Evidence-Based Policymaking Act.
of data and computing environments are central components of the project itself. For example, the LIGO collaboration, mentioned earlier for its use of Jupyter notebooks, has an elaborate data management plan that is updated as the project continues (Anderson and Williams, 2017). The plan explains the overall approach to data stewardship using an Open Archival Information System model, delineates roles and responsibilities for various tasks, and includes provisions for archiving and preserving digital artifacts for LIGO’s users, including the public. Funding comes through central grants to LIGO, as well as through grants to participating investigators and their institutions. Although the trend toward research funders requiring and supporting data sharing is clear, many funders do not currently require data management plans that protect the coherence and completeness of data and objects that are part of a scholarly record.
Implementation Challenges
Efforts to support sharing and persistent access to data, code, and other digital artifacts of research in order to facilitate reproducibility and replicability will need to navigate around several persistent obstacles. For example, to the extent that federal agencies and other research sponsors can harmonize repository requirements and data management plans, it will simplify the tasks associated with operating repositories and perhaps even help to avoid an undue proliferation of repositories. Researchers and research institutions would then find it more straightforward to comply with funder mandates. A consultation or coordinating mechanism among federal agencies and other research sponsors is one possible element toward harmonization and simplification.
Barriers to sharing data and code, such as restrictions on personally identifiable information, national security, and proprietary information, will surely persist. Some research communities in disciplines such as economics rely heavily on nonpublic data and/or code.33 It may be helpful or necessary for such communities to develop alternative mechanisms to verify computational reproducibility. The use of virtual machines is one possible approach, as discussed above.
Finally, as noted in the discussion of repositories, there is intense interest in who controls and owns the tools and infrastructure that researchers use to make digital artifacts available. The relative merits of for-profit companies versus community ownership, as well as open source versus proprietary software, will continue to be important topics of debate as research communities shape their future (National Academies of Sciences, Engineering, and Medicine, 2018).
___________________
33 See https://www.aeaweb.org/research/transparency-reproducibility-credibility-economics.
RECOMMENDATION 6-5: In order to facilitate the transparent sharing and availability of digital artifacts, such as data and code, for its studies, the National Science Foundation (NSF) should
- develop a set of criteria for trusted open repositories to be used by the scientific community for objects of the scholarly record;
- seek to harmonize with other funding agencies the repository criteria and data management plans for scholarly objects;
- endorse or consider creating code and data repositories for long-term archiving and preservation of digital artifacts that support claims made in the scholarly record based on NSF-funded research. These archives could be based at the institutional level or be part of, and harmonized with, the NSF-funded Public Access Repository;
- consider extending NSF’s current data management plan to include other digital artifacts, such as software; and
- work with communities reliant on nonpublic data or code to develop alternative mechanisms for demonstrating reproducibility.
Through these repository criteria, NSF would enable discoverability and standards for digital scholarly objects and discourage an undue proliferation of repositories, perhaps through endorsing or providing one go-to Website that could access NSF-approved repositories.
RECOMMENDATION 6-6: Many stakeholders have a role to play in improving computational reproducibility, including educational institutions, professional societies, researchers, and funders.
- Educational institutions should educate and train students and faculty about computational methods and tools to improve the quality of data and code and to produce reproducible research.
- Professional societies should take responsibility for educating the public and their professional members about the importance and limitations of computational research. Societies have an important role in educating the public about the evolving nature of science and the tools and methods that are used.
- Researchers should collaborate with expert colleagues when their education and training are not adequate to meet the computational requirements of their research.
- In line with its priority for “harnessing the data revolution,” the National Science Foundation (and other funders) should consider funding of activities to promote computational reproducibility.
EFFORTS TO IMPROVE REPLICABILITY
Transparency and complete reporting are key enablers of replicability. No matter how research is conducted, it is essential that other researchers and the public can understand the details of the research study as fully as possible. When researchers “show their work” through detailed methods sections and full transparency about the choices made during the course of research, it introduces a number of advantages. First, transparency allows others to assess the quality of the study and therefore how much weight to give the results. For example, a study that hypothesized an effect on 20 outcomes and found a statistically significant effect on 16 of them would provide stronger evidence for the effectiveness of the intervention than a study that hypothesized an effect on the same 20 outcomes and found a statistically significant effect of the intervention only on 1 outcome. Without transparency, this type of comparison would be impossible. Second, for researchers who may want to replicate a study, transparency means that sufficient details are provided so that the researcher can adhere closely to the original protocol and have the best opportunity to replicate the results. Finally, transparency can serve as an antidote to questionable research practices, such as hypothesizing after results are known or p-hacking, by encouraging researchers to thoroughly track and report the details of the decisions they made and when they made them.
Efforts to foster a culture that values and rewards openness and transparency are taking a number of forms, including guidelines that promote openness, badges and prizes that recognize openness, changes in policies to ensure transparent reporting, new approaches to publishing results, and direct support for replication efforts.
The efforts described here to decrease non-reproducibility and non-replicability have been undertaken by various stakeholders, including journals, funders, educational institutions, and professional societies, as well as researchers themselves. Given the current system’s incentives and roles, publishers and funders can have a strong influence on behavior. For example, funders can make funding contingent on researchers’ following certain practices, and journals can set publication requirements. Professional organizations also have taken steps to improve reproducibility and replicability. They have convened scientists within and across disciplines in order to discuss issues; to develop standards, guidelines, and checklists for ensuring good conduct and reporting of research; and to serve as resources for media in order to improve communication about scientific results. Professional organizations also control some of the incentive structures that can be leveraged to change research practices and norms (e.g., journals, awards, conference presentations).
Openness Guidelines
Professional societies, journals, government organizations, and other stakeholders have worked separately and together on developing guidelines for open sharing. One of the largest efforts was a collaboration among academics, publishers, funders, and industry that began with a 2014 conference at the Lorentz Center in the Netherlands and resulted in the 2016 publication of the FAIR data principles. As noted above, the principles aim to make data findable, accessible, interoperable, and reusable in the hopes that good data management will facilitate scientific discovery (see Wilkinson et al., 2016). Other guidelines include the Transparency and Openness Promotion (TOP) guidelines and those by the Association for Computing Machinery (ACM).34
The TOP guidelines were developed by journals, funders, and societies and published in Science (Nosek et al., 2015). Their goal is to encourage transparency and reproducibility in science; the effort currently has more than 5,000 signatories35:
[The guidelines] include eight modular standards, each with three levels of increasing stringency. Journals select which of the eight transparency standards they wish to implement and select a level of implementation for each. These features provide flexibility for adoption depending on disciplinary variation, but simultaneously establish community standards.
The eight modular standards reflect the discussion throughout this report:
- citation
- data transparency
- analytic methods (code) transparency
- research materials transparency
- design and analysis transparency
- preregistration of studies
- preregistration of analysis plans
- replication
We also note that the definition of “replication” by the TOP administrators is consistent with this committee’s definition.
Several journals have begun requiring researchers to share their data and code. For example, Science implemented a policy in 2011 requiring researchers to make data and code available on request (American Association for the Advancement of Science, 2018):
___________________
34 See https://www.acm.org/publications/task-force-on-data-software-and-reproducibility.
35 See https://cos.io/top.
After publication, all data and materials necessary to understand, assess, and extend the conclusions of the manuscript must be available to any reader of a Science Journal. After publication, all reasonable requests for data, code, or materials must be fulfilled. Any restrictions on the availability of data, code, or materials, including fees and restrictions on original data obtained from other sources must be disclosed to the editors as must any Material Transfer Agreements (MTAs) pertaining to data or materials used or produced in this research, that place constraints on providing these data, code, or materials. Patents (whether applications or awards to the authors or home institutions) related to the work should also be declared.
Journal Requirements, Badges, and Awards
Badges, which recognize and certify open practices, are another way that journals have tried to encourage researchers to share information with the aim of enabling reproducibility. For example, the Center for Open Science has developed three badges—for preregistered studies (see below), open data, and open materials—and at least 34 journals offer one or more of these badges to authors. Initial research indicates that these badges are effective at increasing the rate of data sharing, though not effective in improving other practices, such as code sharing (Kidwell et al., 2016; Rowhani-Farid et al., 2017). ACM has introduced a set of badges for journal articles that certify whether the results have been replicated or reproduced, and whether digital artifacts have been made available or been verified. ACM’s branding structure gives badges in recognition of articles that have passed some level of artifact review (Rous, 2018)36; includes this information in the article metadata, which is searchable in ACM’s digital library; and allows authors to attach code and data to the article’s record. ACM badges were introduced in 2016, and there are now more than 820 articles in the ACM digital library with badges indicating they are accompanied by artifacts (code, data, or both). The IEEE (Institute of Electrical and Electronics Engineers) Xplore Digital Library also assigns reproducibility badges to code or datasets for some articles. Another interesting resource is Papers with Code, a GitHub repository, which is collecting papers that have associated code on GitHub and providing links to the paper, as well as the number of stars on GitHub.37
Awards have also been used to incentivize authors not only to publish reproducible results, but also to do it properly. One example is the Most Reproducible Paper Award, introduced in 2017 by the ACM Special Interest Group on Management of Data.38 There are four criteria for selecting winners:
___________________
36 See https://www.acm.org/publications/policies/artifact-review-badging.
37 See https://github.com/zziz/pwc.
38 See https://sigmod.org/sigmod-awards/sigmod-most-reproducible-paper-award.
- coverage (ideal: all results can be verified)
- ease of reproducibility (ideal: works)
- flexibility (ideal: can change workloads, queries, data and get similar behavior with published results)
- portability (ideal: Linux, Mac, Windows)
Over the past decade, a trend has emerged of journals—including Science, Nature, and PLOS publications—strengthening their data and code sharing policies. Technological advances have changed the journal publishing landscape dramatically, and one advance has been the ability to publish research artifacts connected (and linked) to the manuscript reporting the scientific result. This has led to a push for journals to encourage or require authors to publish the research artifacts necessary for others to attempt to reproduce the results in a manuscript. Other journals do not require this but encourage it, sometimes with extra incentives (Kidwell et al., 2016).
Some journals require authors to make all data underlying their results available at the time of publication (e.g., PLOS publications), while others require authors to make information available on request (e.g., Science). However, as discussed in Chapter 4, authors are not always willing to share, despite these requirements. Some journals now have data editors review and confirm that submissions meet the journal’s data sharing requirements. Others, instead of requiring, give authors the choice to include data and code and provide incentives for them to do so. Information Systems,39 in addition to evaluating the reproducibility of papers, also provides incentives to reviewers: reviewers write, together with the authors of the paper being evaluated, a reproducibility report that is published by the journal. Also, some journals have separated the review of the data and code from the traditional peer review of the overall content of submitted articles.
Introducing Prepublication Checks for Errors and Anomalous Results
Several methods and tools can be used by researchers, peer reviewers, and journals to identify errors in a paper prior to publication. These methods and tools support reproducibility by strengthening the reliability and rigor of results and also deter detrimental research practices such as inappropriate use of statistical analysis as well as data fabrication and falsification. One approach is comparing new results against existing data in the published literature, as in the partnership between the Journal of Chemical and Engineering Data and the Thermodynamics Research Center (TRC) of the National Institute of Standards and Technology. TRC maintains and
___________________
39 See https://www.journals.elsevier.com/information-systems.
curates a number of databases that contain thermophysical and transport properties of pure compounds, binary mixtures, ternary mixtures, and chemical reactions, as well as other data.
Under its partnership with the journal, TRC performs a number of quality checks on the data in articles prior to publication, including identifying anomalous behavior by plotting the new data in different ways and comparison against TRC’s database; confirming that experimental uncertainties are reasonably assessed and reported; and checking that descriptions of sample characterization, method description, and figure and table content are adequate. Papers that fail any of these checks are flagged for follow-up, and the authors can correct the identified mistakes.
Joan Brennecke, editor-in-chief of the journal, reported to the committee that 23 percent of the submissions had “major issues”; of these, around one-third of the anomalies were due to typographical errors or similar mistakes.
Independently reproducing or replicating results before publishing is an effective though time-consuming way to ensure that published results are reproducible or replicable. One journal has undertaken this effort since its inception in 1921; Organic Syntheses, a small journal about the preparation of organic compounds, does not publish research until it has been independently confirmed in the laboratory of a member of the board of editors. In order to facilitate this work, the journal requires authors to provide extensive details about their methods and also expects the authors to have repeated the experiment themselves before submission.40 This type of prepublication check is only feasible for experiments that are relatively simple and inexpensive to reproduce or replicate.
Other approaches to identifying errors prior to publication do not require collaboration with other researchers and could be performed by the authors of papers prior to submission or by journals prior to publication. For example, in psychology and the social sciences, several mathematical tools have been developed to check the statistical data and analyses that use null hypothesis tests, including statcheck and p-checker:
- statcheck independently computes the p-values based on reported statistics, such as the test statistic and degrees of freedom. This tool has also been used to assess the percentage of reported p-values that are inconsistent with their reported data across psychology journals from 1985 to 2013 (Epskamp and Nuijten, 2016; Nuijten et al., 2016).
- p-checker is an app that implements a set of tools developed by various researchers to test whether reported p-values are correct.41
___________________
The tools include p-curve, the R-index, and the test of insufficient variance (Schimmack, 2014; Schönbrodt, 2018; Simonsohn et al., 2014b).
Although these tools have been developed and used in the social science and psychological research communities, they could be used by any researcher or journal to check statistical data in papers using null hypothesis significance tests.
Preregistration of Studies
Preregistration is a specific practice under the broad umbrella of transparency. Confusion over whether an analysis is exploratory or confirmatory can be a source of non-replication, and preregistration can help mitigate this confusion. Specifically, if p-values are interpreted as if the statistical test was planned ahead of time (i.e., confirmatory) when in fact it was exploratory, this will lead to misplaced confidence in the result and an increased likelihood of non-replication. In short, without documentation of which analyses were planned and which were exploratory, p-values (and some other inferential statistics) are easily misinterpreted. One proposed solution is registering the analysis plan before the research is conducted, or at least before the data are examined. This practice goes by different names in different fields, including “pre-analysis plan,” “preregistration,” and “trial registration.” These plans include a precise description of the research design, the statistical analysis that will test the key hypothesis or research question, and the team’s plan for specific analytic decisions (e.g., how sample size will be determined, how outliers will be identified and treated, when and how to transform variables).
Preregistration has several potential advantages (Nosek et al., 2018). First, when done correctly, it makes the researchers’ plans transparent for others to verify. Any deviations from the specified plan would be detectable by others, and the scientific community can decide whether these deviations warrant interpreting the evidence as more exploratory than confirmatory. Second, when done correctly, preregistration improves interpretability of any statistical tests and, when relevant, can ensure that a single, predetermined statistical hypothesis has been carried out and can be regarded as a legitimate test. In addition, the error rate among studies that are preregistered correctly would be controlled (i.e., the rate of false positives when the null hypothesis is true should be equal to alpha, the significance threshold). In other words, preregistration would allow researchers to achieve the stated goal of frequentist hypothesis testing—namely, error control. Third,
by documenting the confirmatory part of their research plan, researchers can expand their design and data collection to simultaneously gather exploratory evidence about new research questions. Without preregistration, a researcher who had an a priori hypothesis but also wanted to collect and analyze data for exploratory purposes in the same study would be indistinguishable from one who had no plan and simply conducted many different tests in a single study. With preregistration, researchers can document their planned hypothesis test and also collect and analyze data for exploratory analyses, and the scientific community can verify that the planned tests were indeed planned. Fourth, preregistration lessens overconfidence in serendipitous results and allows the scientific community to present them with appropriate caveats and take the necessary steps to confirm them.
A common misconception is that preregistration restricts researchers to conducting only the analysis or analyses that were specified in the registered plan. To the contrary, both exploratory and confirmatory analyses can be conducted and reported; preregistration simply allows the scientific community to distinguish between analyses that were prespecified and those that were not and calibrate their confidence accordingly (Simmons, 2018). These new or revised hypotheses would then be subject to new tests conducted with independent data and prespecified analysis plans.
Despite these potential advantages, several concerns about preregistration have been raised (Shiffrin et al., 2018; Morey, 2019). Perhaps the biggest concern is that a preregistration requirement would put an undue burden on researchers without clear evidence that preregistration will actually help lower the instances of false positives. Another concern is that preregistration will change the nature of what researchers study by encouraging them to preferentially test easily confirmable hypotheses and discouraging the kinds of open-ended exploration that can yield important and unanticipated scientific advances (Goldin-Meadow, 2016; Kupferschmidt, 2018). Finally, preregistration is sometimes presented as a proxy for quality research; however, poor-quality ideas, methods, and analyses can be preregistered just like high-quality research.
In a survey of more than 2,000 psychologists, Buttliere and Wicherts (2018) found that preregistration had the lowest support among 11 proposed reforms in psychological science. This lack of acceptance of preregistration may be due, at least in part, to the fact that its effectiveness in changing research practices and improving replication rates is unknown. It could also be due to the fact that tools for preregistering studies have not been around as long as tools for calculating effect sizes or statistical power, so norms surrounding preregistration have had less time to develop.
Encouraging the Publication of All Results
As discussed in Chapter 5, certain results are published over others for a number of reasons. Novel, seemingly consequential, and eye-catching results are favored over replication attempts, null results, or incremental results. Encouraging the publication of null or incremental results will advance replicability by increasing the amount of information available on a scientific topic and reducing the bias favoring the publication of positive effects (Kaplan and Irvin, 2015). The publication of replication studies encourages researchers to expend more effort on replication.
Some new journals stress evaluation of manuscripts based only on quality or rigor, rather than on the newsworthiness or potential impact of the results (e.g., PLOS publications, Collabra: Psychology). In addition to these new specialized journals, some traditional journals have advertised that they welcome submissions of replication studies and studies reporting negative results. With an ever-increasing number of scientific journals, specialized outlets may appear for all kinds of studies (e.g., null results, replications, and methodologically limited studies, as well as groundbreaking results). Some believe that having such outlets will help redress the problem of publication bias, while others doubt the viability of outlets that are likely to be seen as less prestigious. Having outlets for informative but undervalued work does not necessarily mean that researchers will put time and effort into publishing such work, particularly if the incentives do not encourage it (e.g., if the journals that publish such work are not well respected). According to this view, it is important that the same journals that publish original, significant, and eye-catching results also consider publishing important and rigorous work that does not have these features. In the meantime, preprint servers (e.g., arXiv, BioRxiv, PsyArXiv, SocArXiv) make it possible for researchers to post papers whether or not they have been accepted by a scientific journal. Such posting is likely to help the scientific community incorporate the results of a broader range of studies when evaluating the evidence for or against a scientific claim (e.g., when undertaking research syntheses).
The publication of previously unpublishable results is a rather new phenomenon; it will be useful to see if journals that have invited such submissions follow through with publishing such studies, and if so, whether publishing such studies will have an impact (positive or negative) on the journals’ reputations. More importantly, will the availability of outlets lead to more submissions of these types of studies? It is an open question as to what extent the dearth of negative results and replication studies is due to journals’ selection criteria. If more journals and more prestigious journals become more open to publishing these types of studies, a more balanced and realistic literature may result. Journals’ reluctance to publish negative
results and non-groundbreaking studies reflects the value that is currently placed on these types of research by much of the scientific community, which may value novel and eye-catching results over rigor and reliability. While some traditional journals have advertised their willingness to publish negative results and replication studies, these submissions may not be given full consideration in the review process: authors may be subjected to harsher evaluation than if they were to present novel, original, positive results.
A growing number of journals offer the option for submissions to be reviewed on the basis of the proposed research question, method, and proposed analyses, without a reviewer knowing the results.42 The principal argument for this type of approach is to separate the outcome of the study from the evaluation of the importance of the research question and the quality of the research design. The idea is that if the study is testing an important question, and testing it well, then the results should be worthy of publication regardless of the outcome of the study. This approach would not apply to all studies. For example, in some studies the value of the study depends on the outcome, such as “moon-shot” studies testing a very unlikely hypothesis that would be a breakthrough if true, but unremarkable if false.
Two primary ways to conduct results-blind reviewing are available. The first allows authors to submit a version of the manuscript with the results obscured. That is, the results are known to the authors, but they omit the relevant sections from the submitted manuscript so that reviewers must evaluate the submission without knowing the results. Another version of this approach is to invite submissions of manuscripts that propose studies that have not been carried out yet, known as a “registered report” (Chambers et al., 2014). These submissions would include all parts of a manuscript that can be written before data are collected, including background literature and rationale for the research question, proposed methods, and planned analyses. This submission is peer reviewed and the editor can request changes or reject the submission. If accepted, the authors carry out the research and submit a final manuscript with the results, which is again reviewed to ensure that the authors followed the proposed protocol.
No consensus has developed about whether and when the outcome of a study should be a basis for evaluating the publication-worthiness of a paper. Proponents of results-blind review argue that while there are certainly circumstances under which knowing the results should affect the interpretation of the quality of a study (e.g., when the results bring to light error or bias in the study design or analysis), in some circumstances knowing the results could introduce bias into the peer review process. Proponents argue that if more journals offer these options, a more balanced literature would
___________________
42 See http://cos.io/rr.
be produced, and there would therefore be more accurate and replicable conclusions both for individual studies and in meta-analyses. Others are more cautious, arguing that requiring registered reports would be counterproductive, by eliminating the ability of researchers to draw valid inferences that emerge during the study.43 It can also be argued that authors are capable of self-publishing all of their own results in order to avoid publication bias (e.g., by posting them on preprint servers), and that it is reasonable for journals to evaluate results as part of the peer review process.
Additional Journal Initiatives
A number of journals have undertaken initiatives directed at improving reproducibility and replicability. One such example is the Psychology and Cognitive Neuroscience section of Royal Society Open Science, which is committed to publishing replications of empirical research particularly if published within its journal.44 There are new journals that print only negative results, such as New Negatives in Plant Science, as well as efforts by other journals to highlight negative results and failures to replicate, such as PLOS’s “missing pieces” collection of negative, null, and inconclusive results. IEEE Access is an open access journal that publishes negative results, in addition to other articles on topics that do not fit into its traditional journals.45 Some journals have created specific protocols for conducting and publishing replication studies. For example, Advances in Methods and Practices in Psychological Science has a new article type called registered replication reports, which involves multiple labs that all use the same protocols in an attempt to replicate a result. These studies are often conducted in collaboration with the original study authors, and the reports are published regardless of the results. Some journals rely on voluntary badge systems; others require authors to affirm that they have followed certain practices or share their data or code as a requisite for publication.
RECOMMENDATION 6-7: Journals and scientific societies requesting submissions for conferences should disclose their policies relevant to achieving reproducibility and replicability. The strength of the claims made in a journal article or conference submission should reflect the reproducibility and replicability standards to which an article is held, with stronger claims reserved for higher expected levels of reproducibility and replicability. Journals and conference organizers are encouraged to:
___________________
43 See https://www.insidehighered.com/news/2018/02/08/two-journals-experiment-registeredreports-agreeing-publish-articles-based-their.
44 See https://royalsocietypublishing.org/rsos/replication-studies.
45 See https://ieeeaccess.ieee.org/frequently-asked-questions.
- set and implement desired standards of reproducibility and replicability and make this one of their priorities, such as deciding which level they wish to achieve for each Transparency and Openness Promotion guideline and working toward that goal;
- adopt policies to reduce the likelihood of non-replicability, such as considering incentives or requirements for research materials transparency, design, and analysis plan transparency, enhanced review of statistical methods, study or analysis plan preregistration, and replication studies; and
- require as a review criterion that all research reports include a thoughtful discussion of the uncertainty in measurements and conclusions.
Research Funder Efforts to Encourage Replicability
Funders can directly influence how researchers conduct and report studies. By making funding contingent on researchers’ following specific requirements, funders can make certain practices mandatory for large numbers of researchers. For example, a funding agency or philanthropic organization could require that grantees follow certain guidelines for design, conduct, and reporting or could require grantees to preregister hypotheses or publish null results.
Some funders are already taking these types of steps. For example, in 2012, the National Institute of Neurological Disorders and Stroke convened a meeting of stakeholders and published a call for reporting standards, asking researchers to report on blinding, randomization, sample size estimation, and data handling. In 2014, NIH gathered a group of journal editors to discuss reproducibility and replicability. The group developed guidelines for reporting preclinical research that cover statistical analysis, data and material sharing, consideration of refutations, and best practices. About 135 journals have signed on to the guidelines thus far.
Also in 2014, NIH started a process to update application and review language for the research that it funds (National Institutes of Health, 2018d). After collecting input from the community, NIH developed a policy called “enhancing reproducibility through rigor and transparency.” (NIH uses the word “reproducibility” as a broad term to refer to both reproducibility and replicability.) The policy is aimed specifically at rigor and transparency in the hopes that improvements in these areas will result in improved replicability in the long run (see Box 6-2).
In addition to funding requirements, funders can also choose to directly fund reproduction or replication attempts or research syntheses. Replications are sometimes the best and most efficient way to confirm a result so
that decisions can be made and further research can move forward. However, as we note throughout this report, it can be costly and time consuming to replicate previous research, and directing resources at replication means that those resources cannot be used for discovery (refer to Box 5-2, Chapter 5). Given the magnitude of scientific research today—with more than 2 million articles published in 2016 alone—attempting to replicate every study would be daunting and unwise. However, there is undoubtedly a need to replicate some studies in order to confirm or correct the results.
The tension between discovery of new knowledge and confirmation and repudiation of existing knowledge led the committee to develop criteria for investment in replications. These criteria could be used by funders to determine when to fund replication efforts, by journals to determine when to publish replication studies, or by researchers to decide their own research priorities.
Similar criteria have been developed by other bodies (see the example in Table 6-1). The Netherlands Organisation for Scientific Research (NWO) announced in 2016 that it would commit $3.3 million over 3 years for
TABLE 6-1 Assessment of the Desirability of Replication Studies
| Criteria | The desirability of a replication study: |
|---|---|
| Knowledge |
|
| Impact |
|
| Cost |
|
| Alternatives |
|
SOURCE: Royal Netherlands Academy of Arts and Sciences (2018, Table 4).
Dutch scientists who want to reproduce or replicate studies. NWO (2016) has chosen to prioritize funding reproduction and replication in areas of study called “cornerstone research,” defined as research that46
- is cited often,
- has far-reaching consequences for further research,
- plays a major role in policy making,
- is heavily referred to in student textbooks, and
- has received media attention and thus had an impact on public debate.
The Global Young Academy (2018) similarly states that replications may be more necessary “the more applied the research is, or the closer the finding is towards short-term real-life implementation, where the consequence of falsifying data has a greater risk of physical and environmental harm.”
The tradeoff between the resources allocated to research for discovery of new knowledge and to research for replication to confirm or repudiate previous knowledge deserves thoughtful consideration.
Developing Effective Funder Mandates
Policies and mandates can play an important role, but their effectiveness depends on whether they are clear, easy to follow, and harmonized across funders and publishers. Recently, the Public Access Working Group of the Association of American Universities and the Association of Public and Land-grant Universities (2017) released a report with recommendations for federal agencies and research institutions on how they should work individually and collectively to improve the effectiveness of public access policies. To the extent that a harmonized system of digital identifiers for investigators, projects, and outputs can be implemented, the costs and burden of compliance can be reduced, and the process of compliance monitoring can be largely automated. This development would counter the current problem of widespread noncompliance with some open science mandates.
Finally, the nature of data, code, and other digital objects used or generated by different disciplines varies widely. Policies that encourage research communities to define their own requirements and guidelines in order to meet sharing and transparency goals might be helpful in avoiding problems that could arise from the imposition of one-size-fits-all approaches.
The Association of American Universities and the Association of Public and Land-grant Universities (2017) report provides useful guidance for
___________________
46 See https://www.nwo.nl/en/news-and-events/news/2016/nwo-makes-3-million-availablefor-replication-studies-pilot.html.
striking the right balance between harmonization and specificity needed in some aspects of open science policies, and the flexibility that is needed in other areas.
RECOMMENDATION 6-8: Many considerations enter into decisions about what types of scientific studies to fund, including striking a balance between exploratory and confirmatory research. If private or public funders choose to invest in initiatives on reproducibility and replication, two areas may benefit from additional funding:
- education and training initiatives to ensure that researchers have the knowledge, skills, and tools needed to conduct research in ways that adhere to the highest scientific standards; describe methods clearly, specifically, and completely; and express accurately and appropriately the uncertainty involved in the research; and
- reviews of published work, such as testing the reproducibility of published research, conducting rigorous replication studies, and publishing sound critical commentaries.
RECOMMENDATION 6-9: Funders should require a thoughtful discussion in grant applications of how uncertainties will be evaluated, along with any relevant issues regarding replicability and computational reproducibility. Funders should introduce review of reproducibility and replicability guidelines and activities into their merit-review criteria, as a low-cost way to enhance both.
RECOMMENDATION 6-10: When funders, researchers, and other stakeholders are considering whether and where to direct resources for replication studies, they should consider the following criteria:
- The scientific results are important for individual decision making or for policy decisions.
- The results have the potential to make a large contribution to basic scientific knowledge.
- The original result is particularly surprising, that is, it is unexpected in light of previous evidence and knowledge.
- There is controversy about the topic.
- There was potential bias in the original investigation, due, for example, to the source of funding.
- There was a weakness or flaw in the design, methods, or analysis of the original study.
- The cost of a replication is offset by the potential value in reaffirming the original results.
- Future expensive and important studies will build on the original scientific results.
This page intentionally left blank.






































