To take advantage of recent scientific advances in genetics that may lead to improved and precision medicine treatments for psychiatric disorders, the field must systematically and intentionally expand the diversity of large datasets noted Bill Martin in Chapter 2. Such datasets will enable the creation of generalizable, actionable insights applicable to all populations, he said. Yet, currently existing large genomic datasets reflect a distinct European bias, said Sarah Tishkoff, the David and Lyn Silfen University Professor in Genetics and Biology at the University of Pennsylvania. In an examination of genome-wide association studies (GWAS), Tishkoff and colleagues showed that 80 percent of the individuals sampled were of European ancestry, and the same European bias is also reflected in other genomic datasets such as gnomAD1 and GTEx,2 she said (Sirugo et al., 2019). Alicia Martin, a population and statistical geneticist at the Analytic and Translational Genetics Unit at Massachusetts General Hospital and the Broad Institute, noted that this is far out of step with the global population census, where people with European ancestry comprise only about 16 percent of the total, as illustrated in Figure 3-1.
“This is really problematic because it impedes our ability to fully understand the genetic factors that influence human disease and may exacerbate health inequalities,” said Tishkoff. As a result, she said, the translation of genetic research into clinical practice or public health policy may be incomplete or wrong. For example, polygenic risk scores from European-based studies can lead to inaccurate estimation of the significance of pathogenic variants and an individual’s or population’s genetic risk and may hinder the development of appropriate interventions. Polygenic risk scores are constructed using GWAS data to analyze the associations between particu-
___________________
1 To learn more about gnomAD, go to https://gnomad.broadinstitute.org (accessed December 10, 2021).
2 To learn more about GTEx, go to https://gtexportal.org/home (accessed December 10, 2021).
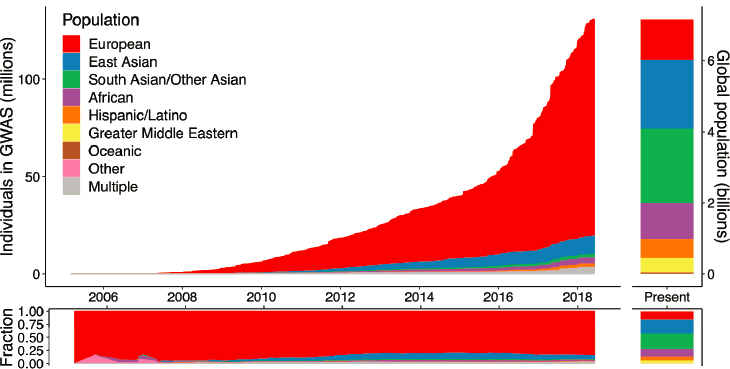
SOURCES: Presented by Alicia Martin, October 5, 2021; Martin et al., 2019.
lar genetic variants and disease outcome and then aggregating the effects of multiple genes. A. Martin added that while polygenic risk scores have tremendous promise, “Right now I think they are more likely to exacerbate health disparities due to these Eurocentric study biases.” She advocated for more diverse GWAS, new methods, increased research capacity, and better communication on culturally sensitive topics related to genetic research.
GENETIC VARIATION ACROSS POPULATIONS AND HEALTH DISPARITIES
The pattern of genetic variation seen in populations today has been shaped by evolutionary history, beginning in Africa where fossils of anatomically modern humans have been dated to about 300,000 years ago, said Tishkoff. Approximately 50,000 to 80,000 years ago, small groups of people migrated out of Africa, giving rise to population groups around the globe. Genetic variation among these groups occurred as a result of migration events, adaptation to different environments, and new spontaneous mutations, said Tishkoff. Yet, even with this variation, most of the genome is shared across populations (as shown in Figure 3-2), she said. About 85 percent of the genetic variation is within populations and only about 10 to 15 percent is between populations (Barbujani et al., 1997; Jorde et al., 2000; Lewontin, 1972).
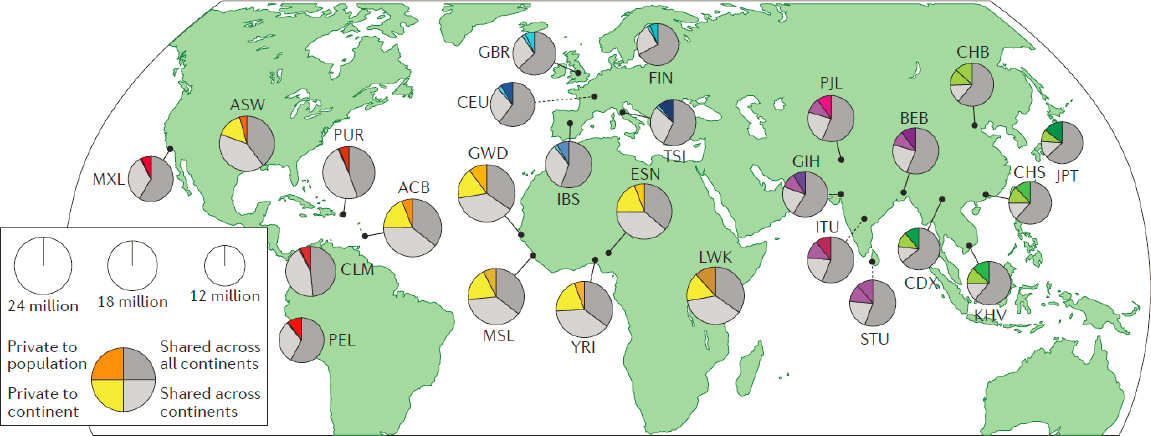
NOTE: ACB = Barbadian; ASW = African Americans in southwest USA; BEB = Bengali; CDX = Dai Chinese; CEU = Utah residents with northern and western European ancestry; CHB = Han Chinese; CHS = southern Han Chinese; CLM = Colombian; ESN = Esan; FIN = Finnish; GBR = British; GIH = Gujarati; GWD = Gambian; IBS = Spanish; ITU = Telugu; JPT = Japanese; KHV = Kinh Vietnamese; LWK = Luhya; MSL = Mende; MXL = Mexican American; PEL = Peruvian; PJL = Punjabi; PUR = Puerto Rican; STU = Tamil; TSI = Tuscan; YRI = Yoruba.
SOURCES: Presented by Sarah Tishkoff, October 5, 2021; Hindorff et al., 2018.
Tishkoff added that health disparities exist for many diseases. For example, individuals with African American ancestry have higher rates of hypertension and those with Native American ancestry have higher rates of type 2 diabetes, she said. Although sociodemographic factors undoubtedly contribute to these disparities, Tishkoff said that underlying genetic factors are believed to be important as well.
Tishkoff and colleagues have delved further into patterns of genetic diversity by examining microsatellite variation in African, African American, and non-African populations (Tishkoff et al., 2009). They showed high levels of genetic diversity within and between populations across Africa, which again reflect evolutionary and demographic history as well as adaptation to diverse environments and diets.
Among African Americans, Tishkoff and colleagues observed predominantly West African ancestry, with an average of about 20 percent European ancestry. The importance of these variations can be seen when looking at specific genomic regions, she said. For example, if a gene involved in drug metabolism shows variations in populations around the world, an individual who self-identifies as African American may have genes from their European ancestry at that particular position, and this could lead to them getting an ineffective treatment. “It emphasizes the need to have knowledge of genome-level variation to enhance precision medicine,” said Tishkoff.
Tishkoff added that natural selection, which occurs as humans adapt to diverse environments, is one important evolutionary force. For example, sickle cell anemia is caused by a mutation that leads to the sickling of red blood cells. People who are homozygous for this mutation will likely die by age 5 without treatment, said Tishkoff. But people who are heterozygous are protected from malaria. Thus, the sickle mutation has risen to a high frequency in regions where malaria is common, she said.
IMPROVING THE PREDICTIVE VALUE OF POLYGENIC RISK SCORES IN PSYCHIATRY
Objective biomarkers are critically needed, but are lacking in psychiatry, in part due to the inaccessibility of brain tissue, said A. Martin. Genetic markers offer a promising solution both as biomarkers and to provide insight into disease mechanisms, she said, because they are straightforward and reproducible to measure. Thus, for example, in recent years genetic studies have shown the importance of synaptic pruning overdrive in schizophrenia (Sekar et al., 2016). Moreover, many psychiatric disorders are highly heritable.
Gene markers are also valuable in assessing individual risk of disease and enabling risk stratification through the use of polygenic risk scores, added A. Martin. One major challenge, she said, is that many genetic variants
have small effect sizes that may contribute to the disease. Schizophrenia, for example, is incredibly polygenic, with nearly 300 genome-wide significant loci identified in the most recent GWAS. “Most of these are non-coding and we don’t know which of these variants are causal, so we have an unknown number of needles and an enormous haystack,” she said.
In addition, polygenic risk scores for patient and non-patient populations often overlap, as shown in Figure 3-3, said A. Martin. Healthy individuals tend to have lower risk because they have fewer genetic risk variants, while those with disease may have higher risk because they have more genetic risk variants. She said looking at the tails of the polygenic risk score distribution can be especially important. Those individuals at the lowest end of the distribution are likely to be protected for a given disease, while those at the highest end are much more likely to have a disease, she added.
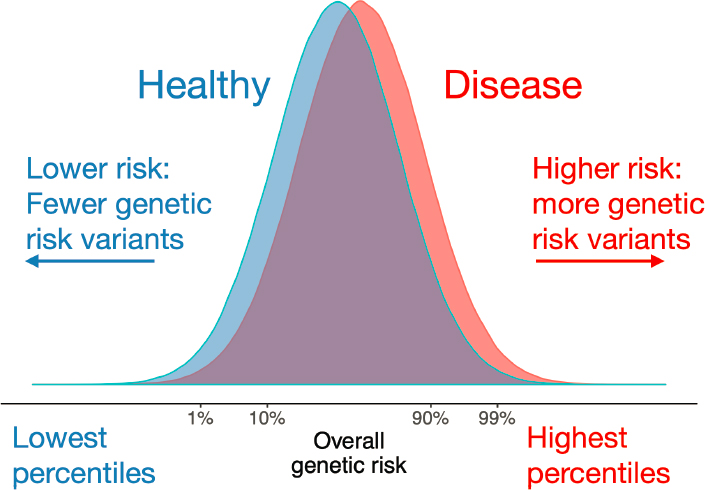
SOURCES: Presented by Alicia Martin, October 5, 2021; Martin et al., 2019.
ADDRESSING THE LACK OF DIVERSITY IN EXISTING GENOMIC DATASETS
As mentioned earlier, genetics has a massive diversity problem, which has become even more pronounced in recent years, said A. Martin. The importance of this lies in the fact that to understand the genetic basis of disease, genetic variation in the population tested is essential. “If we have less genetic variation, or a more biased subset of genetic variation, we’re then destined to find fewer genetic associations that are less representative,” she said. The consequences, she said, are “staggering disparities in accuracy.” With an international group of colleagues, A. Martin compared the prediction accuracy for 17 anthropometric and blood panel traits when using European-derived statistics applied to a massive GWAS done primarily in European ancestry populations. They showed that they could predict these traits about twice as accurately in European ancestry populations compared with East Asian ancestry populations, and about four to five times as accurately as in African ancestry populations (Martin et al., 2019). “I can’t think of any other area of medicine, or any other lab test, that works so differentially in terms of accuracy simply because of who your ancestors are and because of no consequence of your health whatsoever,” she said.
Polygenic prediction in schizophrenia using data from a mostly European ancestry cohort has shown a similar disparity in accuracy, performing about six times more accurately in European ancestry populations than African Americans, and about 1.5 times more accurately in European ancestry populations than in Asian populations, said A. Martin.3 If genetic studies could be made more equitable, polygenic risk scores could have promising clinical use. When studies are not diverse, polygenic scores will be inaccurate for the groups that are being disenfranchised in genetic studies; implementing these scores in the clinic would lead to worsening health inequities, she said.
On the other hand, polygenic risk scores could provide substantial benefits. Among the potential uses of polygenic risk scores in schizophrenia, for example, A. Martin mentioned prediction of medication response, readmission rates, and clinical trial stratification. Increasing diversity in genetic studies could also lead to the identification of more novel biology, including biological processes that are disrupted outside of European ancestry populations, and enable scientists to more accurately pinpoint causal variants and identify genetic and environmental determinants of health, she said.
A. Martin also advocated for integration of multimodal information for complex diseases. For example, cardiovascular disease researchers have
___________________
3 Made available in preprint format. To learn more, go to https://www.medrxiv.org/content/10.1101/2020.09.12.20192922v1 (accessed December 10, 2021).
shown that to predict heart disease risk, combining classical risk factors such as smoking, diabetes, family history, body mass index, hypertension, and high cholesterol with a polygenic score performs better than either alone (Inouye et al., 2018). A. Martin added the caveat that DNA is not destiny, and that many risk factors are modifiable with diet and lifestyle changes.
Indeed, Ekemini Riley, president of the Coalition for Aligning Science and managing director of Aligning Science Across Parkinson’s (ASAP), commented that it would be foolhardy to not include environmental considerations, including what we eat, where we live, and what we breathe in, and how those factors epigenetically modify the entire genome. Diverse datasets provide excellent opportunities to examine gene–environment interactions, for example, by studying populations of Asian ancestry in North America compared with Asians in Asia that encounter different exposures related to diverse diets, lifestyles, and immigration history, said Li-San Wang, professor of pathology and laboratory medicine and founding co-director of the Penn Neurodegeneration Genomics Center at the University of Pennsylvania.
A. Martin noted that the National Institute of Mental Health (NIMH) recently funded the Populations Underrepresented in Mental illness Association Studies (PUMAS), which are aimed at addressing disparity in genetic research. The study aims to recruit and harmonize phenotypes across 180,000 participants in Latin America and Africa; sequence 120,000 genomes from patients with schizophrenia or bipolar disorder as well as ancestry-matched controls; share data and results; and distribute findings.4
Improving the Diversity of Samples in Genetic Research on Alzheimer’s Disease
Alzheimer’s disease (AD) is the most common cause of dementia in older adults, leading to the gradual loss of cognitive abilities and memory, said Wang. Recruiting and retaining participants for AD clinical and genetic studies is challenging both because of the burden on participants and their existing cognitive and health issues, said Wang. Echoing a point made earlier in previous discussions, he added that genetic findings for AD are biased toward European ancestry. Figure 3-4 illustrates how this bias is reflected in what is known about the genetics of AD in non-European populations. For example, Wang noted that while the data indicate that common risk variants for late-onset AD show different effect sizes in European versus African ancestral populations, large diverse datasets are needed to increase
___________________
4 For more information, see https://reporter.nih.gov/search/lWh6-xj4aU-vWGCmZSFoqw/project-details/10263270 (accessed January 31, 2022).
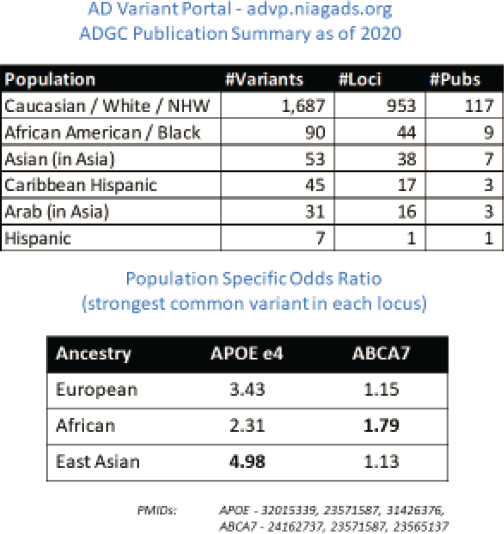
SOURCES: Presented by Li-San Wang, October 5, 2021; Kuksa et al., 2020.
the statistical power for gene discovery and to understand the complete biology of these variants.
The National Institute on Aging has funded several initiatives to increase the sample size and diversity in AD genetic research, said Wang. These included more than 30 AD research centers that have maintained longitudinal cohorts with detailed phenotypes and DNA from diverse populations. More recently, the Alzheimer’s Disease Sequencing Project (ADSP)5 has begun whole-genome sequencing on AD patients. Wang noted that in data released earlier this year from nearly 17,000 completed genomes, almost 30 percent are from underrepresented minorities.
___________________
5 To learn more about ADSP, see https://www.niagads.org/adsp/content/home (accessed December 20, 2021).
Yet, even with these initiatives, a gap exists for Asian Americans and Pacific Islanders, who comprise the fastest growing minority group in the United States. When it comes to individuals of Asian ancestry, Wang said “the cohorts are not there, the investments are not there.” A recent study showed that less than 1 percent of National Institutes of Health (NIH) funding goes to studies among Asian Americans and Pacific Islanders, he said (Doàn et al., 2019). Recruitment challenges in this population include both language and cultural issues and will require building trust with these communities, said Wang. To address these challenges, he and an international group of colleagues from the United States and Canada founded the Asian Cohort for Alzheimer’s Disease (ACAD)6 to recruit Asian participants and leverage available NIH resources for sample management, genotyping capabilities, data management, and data sharing, among others. The ACAD strategy is both community-based and interdisciplinary, and is committed to engaging multiple Asian subpopulations, starting with Chinese, Vietnamese, and Korean, said Wang.
Improving the Diversity of Samples to Understand the Genetic Basis of Parkinson’s Disease
As in other disease areas, lack of diversity in clinical research cohorts has limited understanding of Parkinson’s disease (PD), which affects more than 9 million people globally (Maserejian et al., 2020). The Global Parkinson’s Genetics Program (GP2)7 has partnered with The Michael J. Fox Foundation for Parkinson’s Research to create a rich and diverse dataset and biosamples to drive discoveries in diverse populations, enable the development of human cellular models, and create a framework for examining gene and environment relationships, according to Riley.
GP2 is on pace to genotype more than 150,000 ancestrally diverse volunteers worldwide through multiple strategies, including intentional engagement with other global organizations and capacity building, said Riley. ASAP has committed $35 million for the first 5 years of the project, she said, adding that GP2 partners include the International Parkinson’s Disease Genetics Consortium, the Parkinson’s Foundation’s PD GENEration study,8 the Black and African American Connections to Parkinson’s Disease Study (BLAAC PD),9 and the Latin American Research Consortium on the Genetics
___________________
6 To learn more about ACAD, see https://acadstudy.org (accessed December 10, 2021).
7 To learn more about GP2, see https://gp2.org (accessed December 10, 2021).
8 To learn more about the PD GENEration study, see https://www.parkinson.org/PDGENEration (accessed December 10, 2021).
9 To learn more about BLAAC PD, see https://www.centerwatch.com/clinical-trials/listings/260407/parkinsons-diseaseparkinsons-diseaseparkinsonsparkinson-disease-black-africanamerican-parkinsons-genetic-study/?featured=true (accessed December 10, 2021).
of Parkinson’s Disease (LARGE-PD).10 The map in Figure 3-5 illustrates the locations of samples that have been committed thus far for this project.
Beyond sample gathering, genotyping, and whole-genome sequencing, GP2 has been building global will and capacity, said Riley. Inclusive research and greater participant diversity can be achieved by investing time and demonstrating an interest and willingness “to engage far beyond simply taking samples from a particular group or community,” she said.
INCORPORATING RACE AND ANCESTRY INTO RESEARCH AND CARE
Samples from diverse populations and the potential to encounter extreme phenotypes because of adaptation to different environments could yield important new disease insight and possibly new drug targets, said Tishkoff. However, she noted the importance of ensuring that populations contributing samples benefit from this research, particularly those from low- and middle-income countries that may not even be able to afford new medicines. She also suggested there may be other concerns regarding the wisdom of considering race or ancestry in biomedical research and clinical care.
A. Martin opined that both race and ancestry are critical factors in genetic studies, but cautioned that because socioeconomic and socioenvironmental factors may be unnecessarily coupled with genetics, clarity is needed about the meaning of those labels, why they are relevant for particular models, and the influence of those factors on the study’s findings. This means that public education and careful communication will need to be incorporated into research papers and talking points about the research.
Indeed, added Tishkoff, although race is not a biological construct, it could be important when trying to understand the sociodemographic factors that influence health disparities. A. Martin asked why race is being used in medicine and what is it a proxy for. “Maybe we should just measure those things instead of adding race into the model,” she said.
BARRIERS AND SOLUTIONS TO INCLUDING DIVERSE POPULATIONS IN HUMAN GENOMICS RESEARCH
Workshop speakers identified three major barriers to including diverse populations in human genomics research—time, passion, and historically earned mistrust. “As people start to realize how long it takes to actually gather these cohorts, there’s a bit of trepidation,” said Riley. Moreover, she said, “Going into the community is not an easy task. . . . You need folks
___________________
10 To learn more about LARGE-PD, see https://large-pd.org (accessed December 11, 2021).
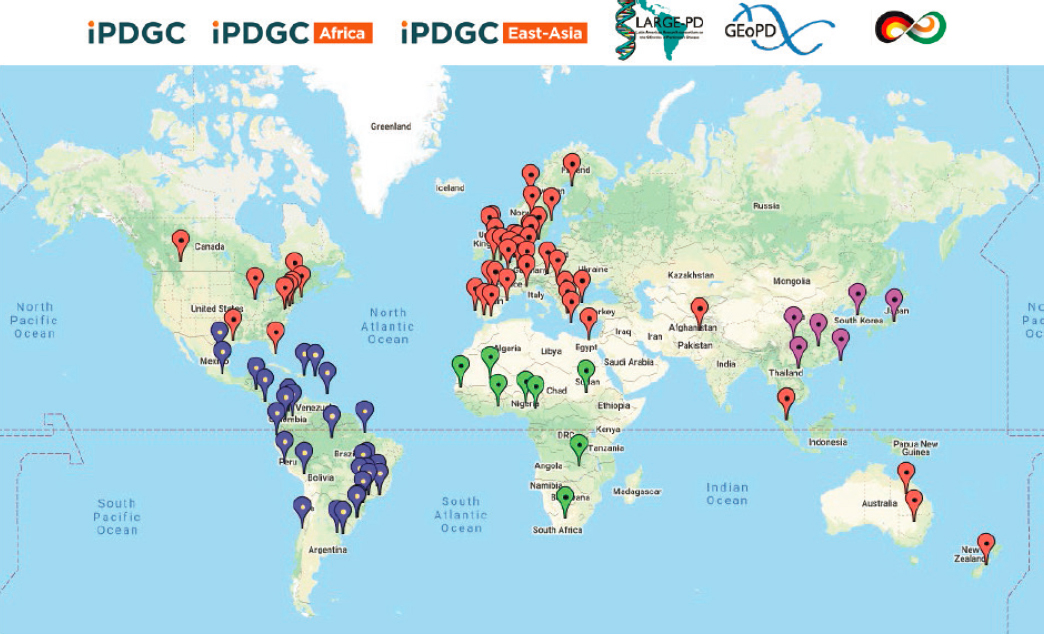
NOTE: For an up-to-date look at committed cohorts and samples, visit the GP2 Cohort Dashboard at https://gp2.org/cohort-dashboard.
SOURCE: Presented by Ekemini Riley, October 5, 2021.
who are really passionate about wanting to see an initiative get off the ground and are willing to put the work in.”
Trust is a critical component in getting these efforts off the ground, noted A. Martin. This requires deep community engagement and leadership that is local and embedded in the community. Moreover, she said, it needs to be earned over time with continual reengagement. This includes informing the community about what has been learned and what the benefits and limitations are of the research. Tishkoff added that gaining trust will require a diverse workforce, including people who look like and represent the desired research participants.
Wang added that investigators and funders also need more awareness of unique concerns in targeted communities. For example, recruiting Asian Americans requires taking into account their immigration history, he said. Studying Asian ancestry in North America compared with Asians in Asia, provides a great opportunity to look at the gene–environment interactions (e.g., how diet or lifestyle affects the disease risk).
Riley and A. Martin added that when going into resource-imbalanced settings, it helps to start by asking people leading initiatives in the specific country what they want and what they need on the ground to make it possible to achieve the research goals. This could include training, equipment, reagents, and mentorship, said A. Martin. “It’s not a one-size-fits-all approach. It’s going to be different for every place.” Moreover, she said, because these needs may change over the course of the project, open lines of communication are critical. Communication also means sharing data and new findings about the disease with these communities, said Wang.
This page intentionally left blank.














